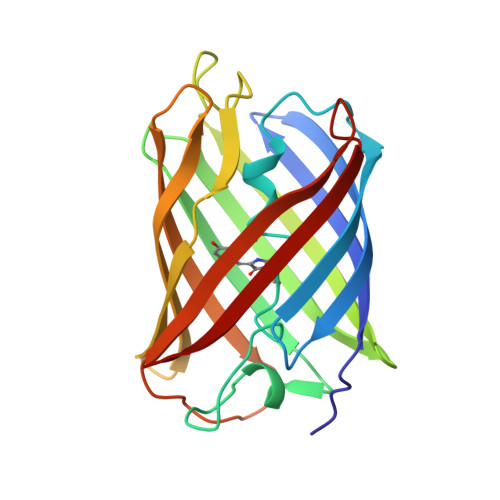
Room Temperature Crystal Structure of the Fast Switching M159T Mutant of the Fluorescent Protein Dronpa.
Kaucikas, M., Fitzpatrick, A., Bryan, E., Struve, A., Henning, R., Kosheleva, I., Srajer, V., Groenhof, G., Van Thor, J.J.(2015) Proteins 83: 397
- PubMed: 25524427
- DOI: https://doi.org/10.1002/prot.24742
- Primary Citation of Related Structures:
4UTS - PubMed Abstract:
The fluorescent protein Dronpa undergoes reversible photoswitching reactions between the bright "on" and dark "off" states via photoisomerization and proton transfer reactions. We report the room temperature crystal structure of the fast switching Met159Thr mutant of Dronpa at 2.0-Å resolution in the bright on state. Structural differences with the wild type include shifted backbone positions of strand β8 containing Thr159 as well as an altered A-C dimer interface involving strands β7, β8, β10, and β11. The Met159Thr mutation increases the cavity volume for the p-hydroxybenzylidene-imidazolinone chromophore as a result of both the side chain difference and the backbone positional differences.
Organizational Affiliation:
Division of Molecular Biosciences, Imperial College London, London, SW7 2AZ, United Kingdom.