Crystal Structure of PVX_084705 with bound PCI32765
Jiang, D.Q., Tempel, W., Loppnau, P., Graslund, S., He, H., Seitova, A., Arrowsmith, C.H., Edwards, A.M., Bountra, C., Hui, R., Hutchinson, A., El Bakkouri, M., Amani, M.To be published.
Experimental Data Snapshot
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cGMP-dependent protein kinase, putative | 847 | Plasmodium vivax Sal-1 | Mutation(s): 0 Gene Names: PVX_084705 | 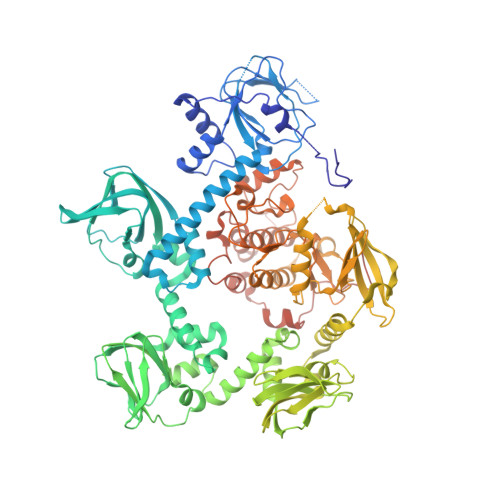 | |
UniProt | |||||
Find proteins for A5K0N4 (Plasmodium vivax (strain Salvador I)) Explore A5K0N4 Go to UniProtKB: A5K0N4 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A5K0N4 | ||||
Sequence AnnotationsExpand | |||||
| |||||
| Ligands 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| 1E8 Query on 1E8 | B [auth A] | 1-{(3R)-3-[4-amino-3-(4-phenoxyphenyl)-1H-pyrazolo[3,4-d]pyrimidin-1-yl]piperidin-1-yl}prop-2-en-1-one C25 H24 N6 O2 XYFPWWZEPKGCCK-GOSISDBHSA-N |  | ||
| UNX Query on UNX | AA [auth A] AB [auth A] BA [auth A] BB [auth A] C [auth A] | UNKNOWN ATOM OR ION X |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 192.556 | α = 90 |
| b = 117.422 | β = 95.26 |
| c = 68.219 | γ = 90 |
| Software Name | Purpose |
|---|---|
| DENZO | data reduction |
| SCALEPACK | data scaling |
| PHASER | phasing |
| PHENIX | refinement |
| PDB_EXTRACT | data extraction |
| APS | data collection |
| HKL-3000 | data reduction |
| HKL-3000 | data scaling |