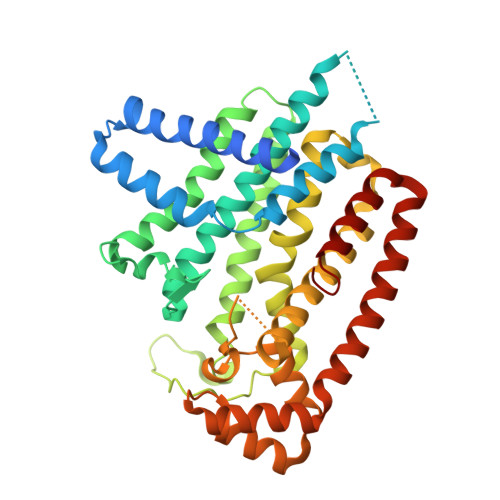
Farnesyl diphosphate synthase inhibitors with unique ligand-binding geometries.
Liu, Y.L., Cao, R., Wang, Y., Oldfield, E.(2015) ACS Med Chem Lett 6: 349-354
- PubMed: 25815158
- DOI: https://doi.org/10.1021/ml500528x
- Primary Citation of Related Structures:
4RXA, 4RXC, 4RXD, 4RXE, 4RYP - PubMed Abstract:
Farnesyl diphosphate synthase (FPPS) is an important drug target for bone resorption, cancer, and some infectious diseases. Here, we report five new structures including two having unique bound ligand geometries. The diamidine inhibitor 7 binds to human FPPS close to the homoallylic (S2) and allosteric (S3) sites and extends into a new site, here called S4. With the bisphosphonate inhibitor 8, two molecules bind to Trypanosoma brucei FPPS, one molecule in the allylic site (S1) and the other close to S2, the first observation of two bisphosphonate molecules bound to FPPS. We also report the structures of apo-FPPS from T. brucei, together with two more bisphosphonate-bound structures (2,9), for purposes of comparison. The diamidine structure is of particular interest because 7 could represent a new lead for lipophilic FPPS inhibitors, while 8 has low micromolar activity against T. brucei, the causative agent of human African trypanosomiasis.
Organizational Affiliation:
Department of Chemistry, University of Illinois at Urbana-Champaign , Urbana, Illinois 61801, United States.