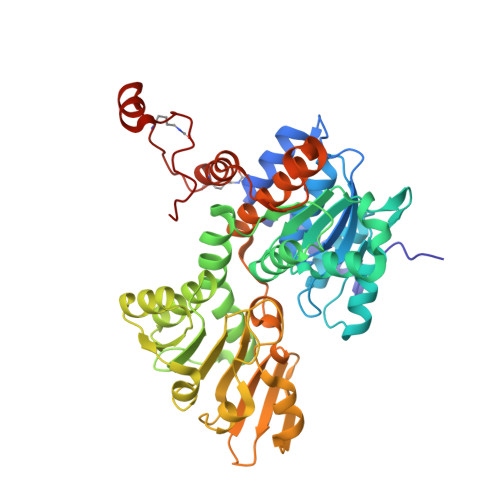
Regulation of s-adenosylhomocysteine hydrolase by lysine acetylation.
Wang, Y., Kavran, J.M., Chen, Z., Karukurichi, K.R., Leahy, D.J., Cole, P.A.(2014) J Biol Chem 289: 31361-31372
- PubMed: 25248746
- DOI: https://doi.org/10.1074/jbc.M114.597153
- Primary Citation of Related Structures:
4PFJ, 4PGF - PubMed Abstract:
S-Adenosylhomocysteine hydrolase (SAHH) is an NAD(+)-dependent tetrameric enzyme that catalyzes the breakdown of S-adenosylhomocysteine to adenosine and homocysteine and is important in cell growth and the regulation of gene expression. Loss of SAHH function can result in global inhibition of cellular methyltransferase enzymes because of high levels of S-adenosylhomocysteine. Prior proteomics studies have identified two SAHH acetylation sites at Lys(401) and Lys(408) but the impact of these post-translational modifications has not yet been determined. Here we use expressed protein ligation to produce semisynthetic SAHH acetylated at Lys(401) and Lys(408) and show that modification of either position negatively impacts the catalytic activity of SAHH. X-ray crystal structures of 408-acetylated SAHH and dually acetylated SAHH have been determined and reveal perturbations in the C-terminal hydrogen bonding patterns, a region of the protein important for NAD(+) binding. These crystal structures along with mutagenesis data suggest that such hydrogen bond perturbations are responsible for SAHH catalytic inhibition by acetylation. These results suggest how increased acetylation of SAHH may globally influence cellular methylation patterns.
Organizational Affiliation:
From the Deptartments of Pharmacology and Molecular Sciences and.