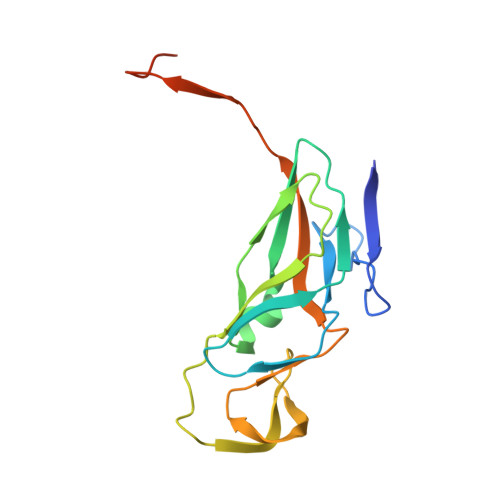
Structure and enzymatic mechanism of a moonlighting dUTPase
Leveles, I., Nemeth, V., Szabo, J.E., Harmat, V., Nyiri, K., Bendes, A.A., Papp-Kadar, V., Zagyva, I., Rona, G., Ozohanics, O., Vekey, K., Toth, J., Vertessy, B.G.(2013) Acta Crystallogr D Biol Crystallogr 69: 2298-2308
- PubMed: 24311572
- DOI: https://doi.org/10.1107/S0907444913021136
- Primary Citation of Related Structures:
4GV8 - PubMed Abstract:
Genome integrity requires well controlled cellular pools of nucleotides. dUTPases are responsible for regulating cellular dUTP levels and providing dUMP for dTTP biosynthesis. In Staphylococcus, phage dUTPases are also suggested to be involved in a moonlighting function regulating the expression of pathogenicity-island genes. Staphylococcal phage trimeric dUTPase sequences include a specific insertion that is not found in other organisms. Here, a 2.1 Å resolution three-dimensional structure of a ϕ11 phage dUTPase trimer with complete localization of the phage-specific insert, which folds into a small β-pleated mini-domain reaching out from the dUTPase core surface, is presented. The insert mini-domains jointly coordinate a single Mg2+ ion per trimer at the entrance to the threefold inner channel. Structural results provide an explanation for the role of Asp95, which is suggested to have functional significance in the moonlighting activity, as the metal-ion-coordinating moiety potentially involved in correct positioning of the insert. Enzyme-kinetics studies of wild-type and mutant constructs show that the insert has no major role in dUTP binding or cleavage and provide a description of the elementary steps (fast binding of substrate and release of products). In conclusion, the structural and kinetic data allow insights into both the phage-specific characteristics and the generally conserved traits of ϕ11 phage dUTPase.
Organizational Affiliation:
Institute of Enzymology, Research Centre for Natural Sciences, Hungarian Academy of Sciences, 29 Karolina Street, 1113 Budapest, Hungary.