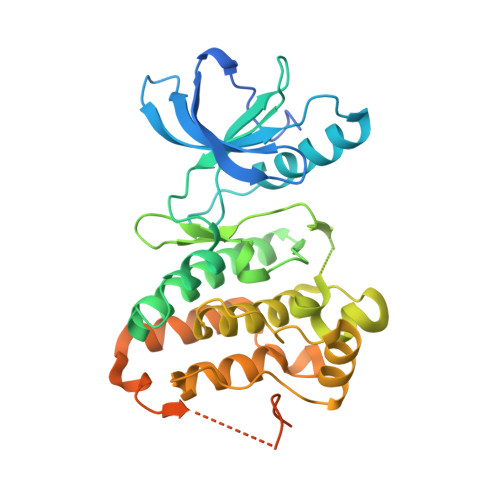
Discovery of a novel chemotype of tyrosine kinase inhibitors by fragment-based docking and molecular dynamics.
Zhao, H., Dong, J., Lafleur, K., Nevado, C., Caflisch, A.(2012) ACS Med Chem Lett 3: 834-838
- PubMed: 24900387
- DOI: https://doi.org/10.1021/ml3001984
- Primary Citation of Related Structures:
4G2F - PubMed Abstract:
We have discovered a novel chemical class of inhibitors of the EphB4 tyrosine kinase by fragment-based high-throughput docking followed by explicit solvent molecular dynamics simulations for assessment of the binding mode. The synthesis of a single derivative (compound 7) of the hit identified in silico has resulted in an improvement of the inhibitory potency in an enzymatic assay from 8.4 μM to 160 nM and a ligand efficiency of 0.39 kcal/mol per non-hydrogen atom. Such remarkable improvement in affinity is due to an additional hydroxyl group involved in two favorable (buried) hydrogen bonds as predicted by molecular dynamics and validated by the crystal structure of the complex with EphA3 solved at 1.7 Å resolution.
Organizational Affiliation:
Department of Biochemistry and Department of Organic Chemistry, University of Zurich , Winterthurerstrasse 190, CH-8057 Zurich, Switzerland.