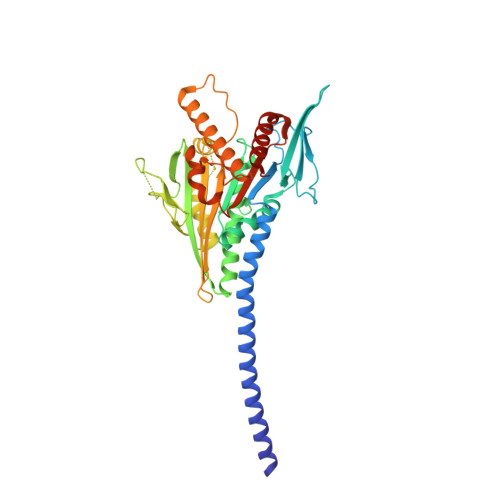
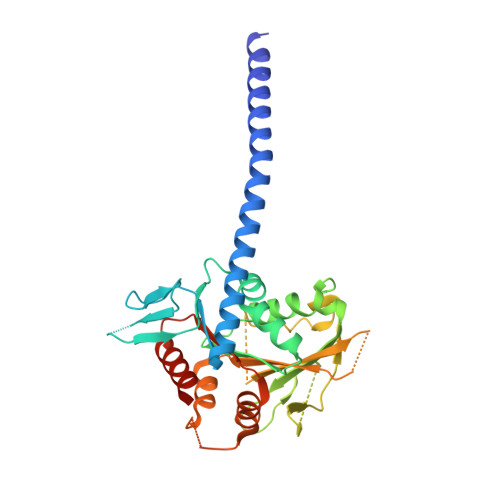
Kar3Vik1, a member of the kinesin-14 superfamily, shows a novel kinesin microtubule binding pattern.
Rank, K.C., Chen, C.J., Cope, J., Porche, K., Hoenger, A., Gilbert, S.P., Rayment, I.(2012) J Cell Biol 197: 957-970
- PubMed: 22734002
- DOI: https://doi.org/10.1083/jcb.201201132
- Primary Citation of Related Structures:
4ETP - PubMed Abstract:
Kinesin-14 motors generate microtubule minus-end-directed force used in mitosis and meiosis. These motors are dimeric and operate with a nonprocessive powerstroke mechanism, but the role of the second head in motility has been unclear. In Saccharomyces cerevisiae, the Kinesin-14 Kar3 forms a heterodimer with either Vik1 or Cik1. Vik1 contains a motor homology domain that retains microtubule binding properties but lacks a nucleotide binding site. In this case, both heads are implicated in motility. Here, we show through structural determination of a C-terminal heterodimeric Kar3Vik1, electron microscopy, equilibrium binding, and motility that at the start of the cycle, Kar3Vik1 binds to or occludes two αβ-tubulin subunits on adjacent protofilaments. The cycle begins as Vik1 collides with the microtubule followed by Kar3 microtubule association and ADP release, thereby destabilizing the Vik1-microtubule interaction and positioning the motor for the start of the powerstroke. The results indicate that head-head communication is mediated through the adjoining coiled coil.
Organizational Affiliation:
Department of Biochemistry, University of Wisconsin, Madison, WI 53706, USA.