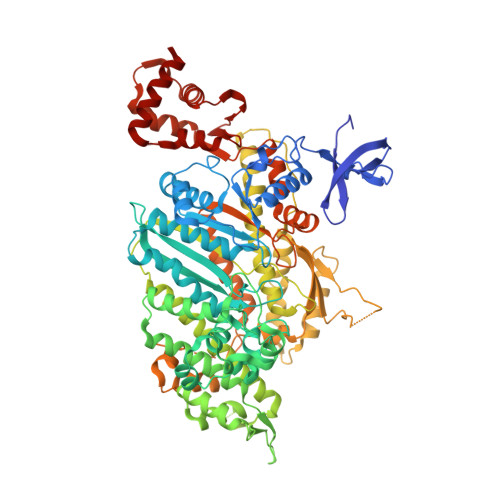
Processive Steps in the Reverse Direction Require Uncoupling of the Lead Head Lever Arm of Myosin VI.
Menetrey, J., Isabet, T., Ropars, V., Mukherjea, M., Pylypenko, O., Liu, X., Perez, J., Vachette, P., Sweeney, H.L., Houdusse, A.M.(2012) Mol Cell 48: 75-86
- PubMed: 22940248
- DOI: https://doi.org/10.1016/j.molcel.2012.07.034
- Primary Citation of Related Structures:
4ANJ, 4E7S, 4E7Z - PubMed Abstract:
Myosin VI is the only known reverse-direction myosin motor. It has an unprecedented means of amplifying movements within the motor involving rearrangements of the converter subdomain at the C terminus of the motor and an unusual lever arm projecting from the converter. While the average step size of a myosin VI dimer is 30-36 nm, the step size is highly variable, presenting a challenge to the lever arm mechanism by which all myosins are thought to move. Herein, we present structures of myosin VI that reveal regions of compliance that allow an uncoupling of the lead head when movement is modeled on actin. The location of the compliance restricts the possible actin binding sites and predicts the observed stepping behavior. The model reveals that myosin VI, unlike plus-end directed myosins, does not use a pure lever arm mechanism, but instead steps with a mechanism analogous to the kinesin neck-linker uncoupling model.
Organizational Affiliation:
Structural Motility, Institut Curie, Centre de Recherche, Paris, France.