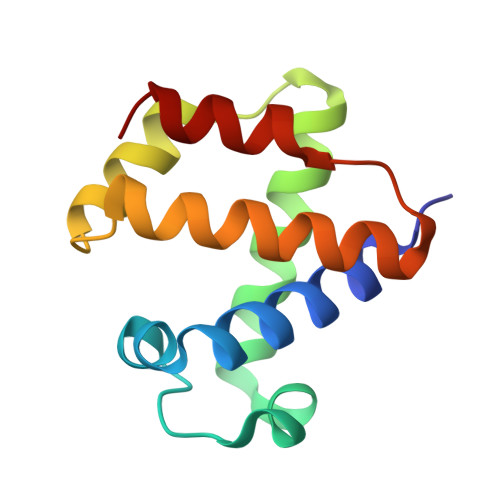
High Resolution Crystal Structures of the Cerebratulus Lacteus Mini-Hb in the Unligated and Carbomonoxy States.
Germani, F., Pesce, A., Venturini, A., Moens, L., Bolognesi, M., Dewilde, S., Nardini, M.(2012) Int J Mol Sci 13: 8025
- PubMed: 22942687
- DOI: https://doi.org/10.3390/ijms13078025
- Primary Citation of Related Structures:
4AVD, 4AVE - PubMed Abstract:
The nerve tissue mini-hemoglobin from Cerebratulus lacteus (CerHb) displays an essential globin fold hosting a protein matrix tunnel held to allow traffic of small ligands to and from the heme. CerHb heme pocket hosts the distal TyrB10/GlnE7 pair, normally linked to low rates of O(2) dissociation and ultra-high O(2) affinity. However, CerHb affinity for O(2) is similar to that of mammalian myoglobins, due to a dynamic equilibrium between high and low affinity states driven by the ability of ThrE11 to orient the TyrB10 OH group relative to the heme ligand. We present here the high resolution crystal structures of CerHb in the unligated and carbomonoxy states. Although CO binds to the heme with an orientation different from the O(2) ligand, the overall binding schemes for CO and O(2) are essentially the same, both ligands being stabilized through a network of hydrogen bonds based on TyrB10, GlnE7, and ThrE11. No dramatic protein structural changes are needed to support binding of the ligands, which can freely reach the heme distal site through the apolar tunnel. A lack of main conformational changes between the heme-unligated and -ligated states grants stability to the folded mini-Hb and is a prerequisite for fast ligand diffusion to/from the heme.
Organizational Affiliation:
Department of Biomedical Sciences, University of Antwerp, Universiteitsplein 1, B-2610 Antwerp, Belgium.