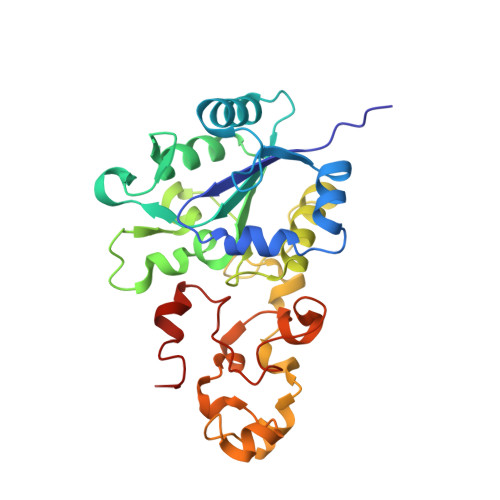
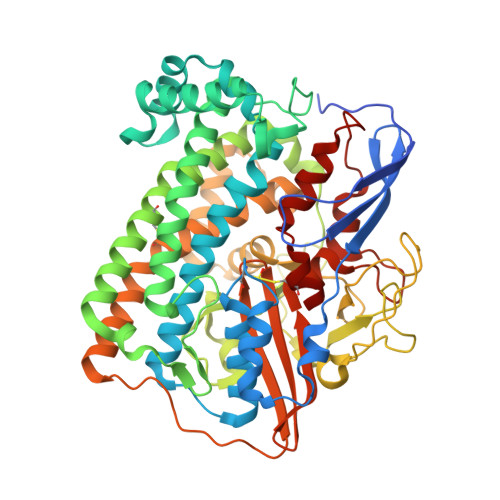
A Threonine Stabilizes the Nic and Nir Catalytic Intermediates of [Nife]-Hydrogenase.
Abou-Hamdan, A., Ceccaldi, P., Lebrette, H., Gutierrez-Sanz, O., Richaud, P., Cournac, L., Guigliarelli, B., De Lacey, A.L., Leger, C., Volbeda, A., Burlat, B., Dementin, S.(2015) J Biol Chem 290: 8550
- PubMed: 25666617
- DOI: https://doi.org/10.1074/jbc.M114.630491
- Primary Citation of Related Structures:
4UCQ, 4UCW, 4UCX - PubMed Abstract:
The heterodimeric [NiFe] hydrogenase from Desulfovibrio fructosovorans catalyzes the reversible oxidation of H2 into protons and electrons. The catalytic intermediates have been attributed to forms of the active site (NiSI, NiR, and NiC) detected using spectroscopic methods under potentiometric but non-catalytic conditions. Here, we produced variants by replacing the conserved Thr-18 residue in the small subunit with Ser, Val, Gln, Gly, or Asp, and we analyzed the effects of these mutations on the kinetic (H2 oxidation, H2 production, and H/D exchange), spectroscopic (IR, EPR), and structural properties of the enzyme. The mutations disrupt the H-bond network in the crystals and have a strong effect on H2 oxidation and H2 production turnover rates. However, the absence of correlation between activity and rate of H/D exchange in the series of variants suggests that the alcoholic group of Thr-18 is not necessarily a proton relay. Instead, the correlation between H2 oxidation and production activity and the detection of the NiC species in reduced samples confirms that NiC is a catalytic intermediate and suggests that Thr-18 is important to stabilize the local protein structure of the active site ensuring fast NiSI-NiC-NiR interconversions during H2 oxidation/production.
Organizational Affiliation:
From the Laboratoire de Bioénergétique et Ingénierie des Protéines, Institut de Microbiologie de la Méditerranée, UMR 7281 Aix-Marseille Université/CNRS, 31 Chemin J. Aiguier, 13402 Marseille Cedex 20, France.