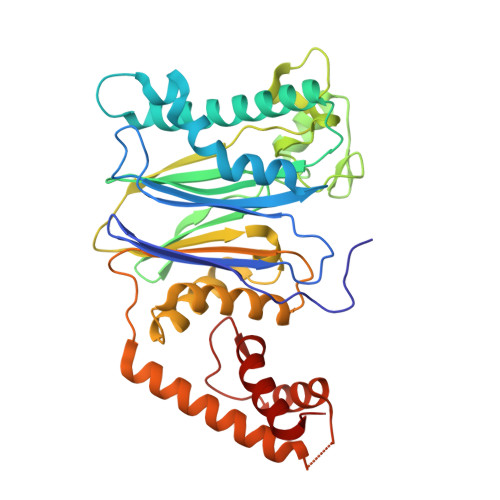
The catalytic role of the M2 metal ion in PP2C alpha
Pan, C., Tang, J.Y., Xu, Y.F., Xiao, P., Liu, H.D., Wang, H.A., Wang, W.B., Meng, F.G., Yu, X., Sun, J.P.(2015) Sci Rep 5: 8560-8560
- PubMed: 25708299
- DOI: https://doi.org/10.1038/srep08560
- Primary Citation of Related Structures:
4RAF, 4RAG - PubMed Abstract:
PP2C family phosphatases (the type 2C family of protein phosphatases; or metal-dependent phosphatase, PPM) constitute an important class of signaling enzymes that regulate many fundamental life activities. All PP2C family members have a conserved binuclear metal ion active center that is essential for their catalysis. However, the catalytic role of each metal ion during catalysis remains elusive. In this study, we discovered that mutations in the structurally buried D38 residue of PP2Cα (PPM1A) redefined the water-mediated hydrogen network in the active site and selectively disrupted M2 metal ion binding. Using the D38A and D38K mutations of PP2Cα as specific tools in combination with enzymology analysis, our results demonstrated that the M2 metal ion determines the rate-limiting step of substrate hydrolysis, participates in dianion substrate binding and stabilizes the leaving group after P-O bond cleavage. The newly characterized catalytic role of the M2 metal ion in this family not only provides insight into how the binuclear metal centers of the PP2C phosphatases are organized for efficient catalysis but also helps increase our understanding of the function and substrate specificity of PP2C family members.
Organizational Affiliation:
1] Key Laboratory Experimental Teratology of the Ministry of Education and Department of Biochemistry and Molecular Biology, Shandong University, School of Medicine, Jinan, Shandong, China [2] Qilu Hospital of Shandong University, Jinan, China.