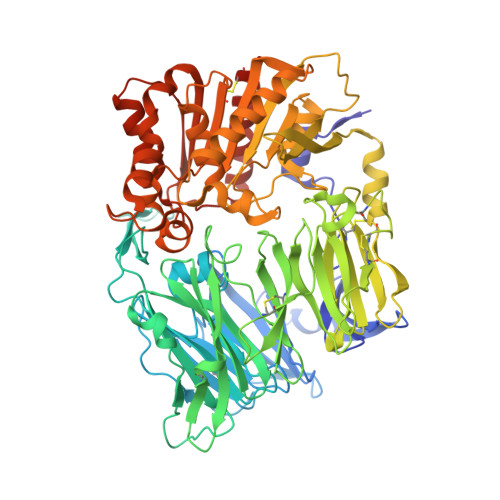
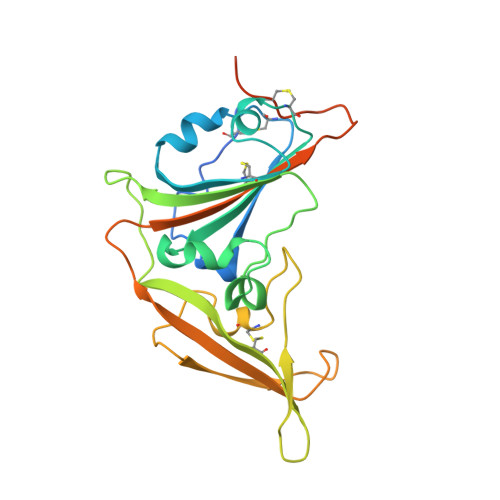
Bat Origins of MERS-CoV Supported by Bat Coronavirus HKU4 Usage of Human Receptor CD26.
Wang, Q.H., Qi, J.X., Yuan, Y., Xuan, Y., Han, P., Wan, Y., Ji, W., Li, Y., Wu, Y., Wang, J., Iwamoto, A., Woo, P.C., Yuen, K.Y., Yan, J., Lu, G.W., Gao, G.F.(2014) Cell Host Microbe 16: 328-337
- PubMed: 25211075
- DOI: https://doi.org/10.1016/j.chom.2014.08.009
- Primary Citation of Related Structures:
4QZV - PubMed Abstract:
The recently reported Middle East respiratory syndrome coronavirus (MERS-CoV) is phylogenetically closely related to the bat coronaviruses (BatCoVs) HKU4 and HKU5. However, the evolutionary pathway of MERS-CoV is still unclear. A receptor binding domain (RBD) in the MERS-CoV envelope-embedded spike protein specifically engages human CD26 (hCD26) to initiate viral entry. The high sequence identity in the viral spike protein prompted us to investigate if HKU4 and HKU5 can recognize hCD26 for cell entry. We found that HKU4-RBD, but not HKU5-RBD, binds to hCD26, and pseudotyped viruses embedding HKU4 spike can infect cells via hCD26 recognition. The structure of the HKU4-RBD/hCD26 complex revealed a hCD26-binding mode similar overall to that observed for MERS-RBD. HKU4-RBD, however, is less adapted to hCD26 than MERS-RBD, explaining its lower affinity for receptor binding. Our findings support a bat origin for MERS-CoV and indicate the need for surveillance of HKU4-related viruses in bats.
Organizational Affiliation:
CAS Key Laboratory of Pathogenic Microbiology and Immunology, Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China.