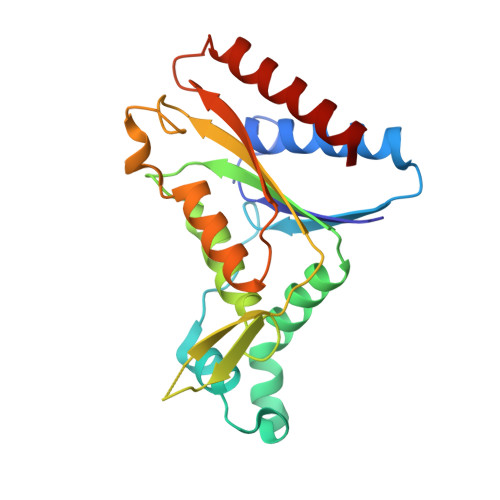
Identification of NAD(P)H Quinone Oxidoreductase Activity in Azoreductases from P. aeruginosa: Azoreductases and NAD(P)H Quinone Oxidoreductases Belong to the Same FMN-Dependent Superfamily of Enzymes.
Ryan, A., Kaplan, E., Nebel, J.C., Polycarpou, E., Crescente, V., Lowe, E., Preston, G.M., Sim, E.(2014) PLoS One 9: e98551-e98551
- PubMed: 24915188
- DOI: https://doi.org/10.1371/journal.pone.0098551
- Primary Citation of Related Structures:
4N65, 4N9Q - PubMed Abstract:
Water soluble quinones are a group of cytotoxic anti-bacterial compounds that are secreted by many species of plants, invertebrates, fungi and bacteria. Studies in a number of species have shown the importance of quinones in response to pathogenic bacteria of the genus Pseudomonas. Two electron reduction is an important mechanism of quinone detoxification as it generates the less toxic quinol. In most organisms this reaction is carried out by a group of flavoenzymes known as NAD(P)H quinone oxidoreductases. Azoreductases have previously been separate from this group, however using azoreductases from Pseudomonas aeruginosa we show that they can rapidly reduce quinones. Azoreductases from the same organism are also shown to have distinct substrate specificity profiles allowing them to reduce a wide range of quinones. The azoreductase family is also shown to be more extensive than originally thought, due to the large sequence divergence amongst its members. As both NAD(P)H quinone oxidoreductases and azoreductases have related reaction mechanisms it is proposed that they form an enzyme superfamily. The ubiquitous and diverse nature of azoreductases alongside their broad substrate specificity, indicates they play a wide role in cellular survival under adverse conditions.
Organizational Affiliation:
Department of Pharmacology, University of Oxford, Oxford, United Kingdom; Faculty of Science, Engineering and Computing, Kingston University, Kingston upon Thames, United Kingdom.