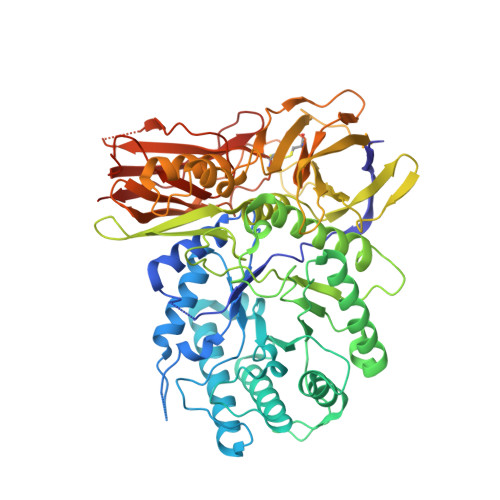
Insights into mucopolysaccharidosis I from the structure and action of alpha-L-iduronidase.
Bie, H., Yin, J., He, X., Kermode, A.R., Goddard-Borger, E.D., Withers, S.G., James, M.N.(2013) Nat Chem Biol 9: 739-745
- PubMed: 24036510
- DOI: https://doi.org/10.1038/nchembio.1357
- Primary Citation of Related Structures:
4KGJ, 4KGL, 4KH2, 4MJ2, 4MJ4 - PubMed Abstract:
Mucopolysaccharidosis type I (MPS I), caused by mutations in the gene encoding α-L-iduronidase (IDUA), is one of approximately 70 genetic disorders collectively known as the lysosomal storage diseases. To gain insight into the basis for MPS I, we crystallized human IDUA produced in an Arabidopsis thaliana cgl mutant. IDUA consists of a TIM barrel domain containing the catalytic site, a β-sandwich domain and a fibronectin-like domain. Structures of IDUA bound to iduronate analogs illustrate the Michaelis complex and reveal a (2,5)B conformation in the glycosyl-enzyme intermediate, which suggest a retaining double displacement reaction involving the nucleophilic Glu299 and the general acid/base Glu182. Unexpectedly, the N-glycan attached to Asn372 interacts with iduronate analogs in the active site and is required for enzymatic activity. Finally, these IDUA structures and biochemical analysis of the disease-relevant P533R mutation have enabled us to correlate the effects of mutations in IDUA to clinical phenotypes.
Organizational Affiliation:
1] Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada. [2] These authors contributed equally to this work.