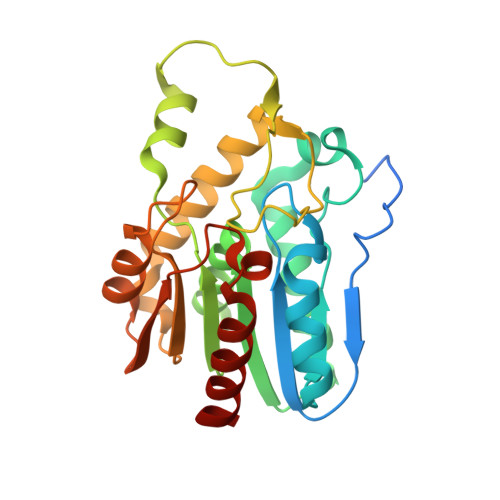
Conformational plasticity and ligand binding of bacterial monoacylglycerol lipase.
Rengachari, S., Aschauer, P., Schittmayer, M., Mayer, N., Gruber, K., Breinbauer, R., Birner-Gruenberger, R., Dreveny, I., Oberer, M.(2013) J Biol Chem 288: 31093-31104
- PubMed: 24014019
- DOI: https://doi.org/10.1074/jbc.M113.491415
- Primary Citation of Related Structures:
4KE6, 4KE7, 4KE8, 4KE9, 4KEA - PubMed Abstract:
Monoacylglycerol lipases (MGLs) play an important role in lipid catabolism across all kingdoms of life by catalyzing the release of free fatty acids from monoacylglycerols. The three-dimensional structures of human and a bacterial MGL were determined only recently as the first members of this lipase family. In addition to the α/β-hydrolase core, they showed unexpected structural similarities even in the cap region. Nevertheless, the structural basis for substrate binding and conformational changes of MGLs is poorly understood. Here, we present a comprehensive study of five crystal structures of MGL from Bacillus sp. H257 in its free form and in complex with different substrate analogs and the natural substrate 1-lauroylglycerol. The occurrence of different conformations reveals a high degree of conformational plasticity of the cap region. We identify a specific residue, Ile-145, that might act as a gatekeeper restricting access to the binding site. Site-directed mutagenesis of Ile-145 leads to significantly reduced hydrolase activity. Bacterial MGLs in complex with 1-lauroylglycerol, myristoyl, palmitoyl, and stearoyl substrate analogs enable identification of the binding sites for the alkyl chain and the glycerol moiety of the natural ligand. They also provide snapshots of the hydrolytic reaction of a bacterial MGL at different stages. The alkyl chains are buried in a hydrophobic tunnel in an extended conformation. Binding of the glycerol moiety is mediated via Glu-156 and water molecules. Analysis of the structural features responsible for cap plasticity and the binding modes of the ligands suggests conservation of these features also in human MGL.
Organizational Affiliation:
From the Institute of Molecular Biosciences, University of Graz, Humboldtstrasse 50/3, A-8010 Graz, Austria.