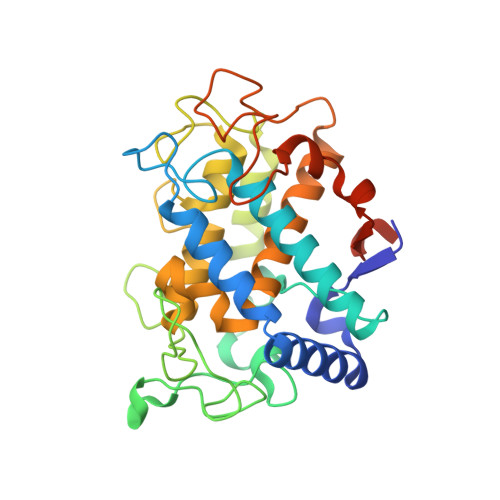
The mechanism of copper uptake by tyrosinase from Bacillus megaterium.
Kanteev, M., Goldfeder, M., Chojnacki, M., Adir, N., Fishman, A.(2013) J Biol Inorg Chem 18: 895-903
- PubMed: 24061559
- DOI: https://doi.org/10.1007/s00775-013-1034-0
- Primary Citation of Related Structures:
4J6T, 4J6U, 4J6V - PubMed Abstract:
Tyrosinase belongs to the type 3 copper enzyme family, containing a dinuclear copper center, CuA and CuB. It is mainly responsible for melanin production in a wide range of organisms. Although copper ions are essential for the activity of tyrosinase, the mechanism of copper uptake is still unclear. We have recently determined the crystal structure of tyrosinase from Bacillus megaterium (TyrBm) and revealed that this enzyme has tighter binding of CuA in comparison with CuB. Investigating copper accumulation in TyrBm, we found that the presence of copper has a more significant effect on the diphenolase activity. By decreasing the concentration of copper, we increased the diphenolase to monophenolase activity ratio twofold. Using a rational design approach, we identified five variants having an impact on copper uptake. We have found that a major role of the highly conserved Asn205 residue is to stabilize the orientation of the His204 imidazole ring in the binding site, thereby promoting the correct coordination of CuB. Further investigation of these variants revealed that Phe197, Met61, and Met184, which are located at the entrance to the binding site, not only play a role in copper uptake, but are also important for enhancing the diphenolase activity. We propose a mechanism of copper accumulation by the enzyme as well as an approach to changing the selectivity of TyrBm towards L-dopa production.
Organizational Affiliation:
Department of Biotechnology and Food Engineering, Technion-Israel Institute of Technology, 32000, Haifa, Israel.