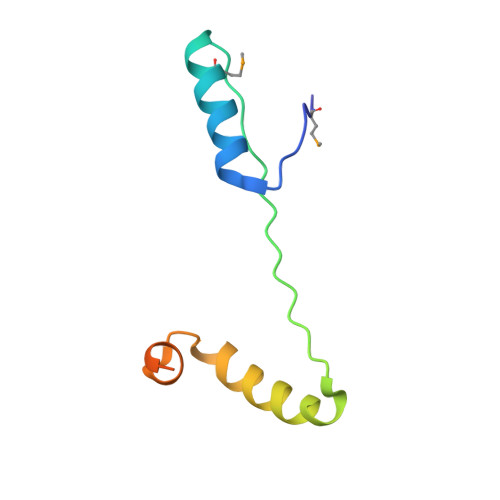
Domain swapping in the cytoplasmic domain of the Escherichia coli rhomboid protease.
Lazareno-Saez, C., Arutyunova, E., Coquelle, N., Lemieux, M.J.(2013) J Mol Biol 425: 1127-1142
- PubMed: 23353827
- DOI: https://doi.org/10.1016/j.jmb.2013.01.019
- Primary Citation of Related Structures:
4HDD - PubMed Abstract:
Rhomboids are membrane-embedded serine proteases that cleave membrane protein substrates. Escherichia coli rhomboid GlpG (ecGlpG) consists of an N-terminal cytoplasmic domain and a membrane domain containing the active site. We determined the crystal structure of the soluble cytoplasmic domain of ecGlpG at 1.35Å resolution and examined whether this domain affected the catalytic activity of the enzyme. The structure revealed that the ecGlpG cytoplasmic domain exists as a dimer with extensive domain swapping between the two monomers. Domain-swapped dimers can be isolated from the full-length protein, suggesting that this is a physiologically relevant structure. An extensive steady-state kinetic analysis of the full-length ecGlpG and its membrane domain using soluble and transmembrane model protein substrates resulted in an unexpected conclusion: removal of the cytoplasmic domain does not alter the catalytic parameters for detergent-solubilized rhomboid for both substrates.
Organizational Affiliation:
Membrane Protein Disease Research Group, Department of Biochemistry, Faculty of Medicine and Dentistry, University of Alberta, Edmonton, Alberta, Canada T6G 2H7.