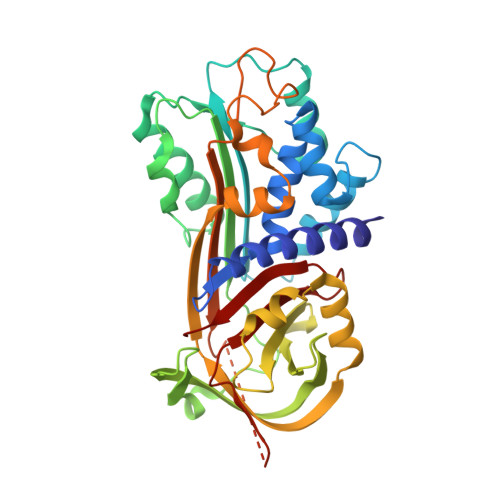
Mechanistic characterization and crystal structure of a small molecule inactivator bound to plasminogen activator inhibitor-1.
Li, S.H., Reinke, A.A., Sanders, K.L., Emal, C.D., Whisstock, J.C., Stuckey, J.A., Lawrence, D.A.(2013) Proc Natl Acad Sci U S A 110: E4941-E4949
- PubMed: 24297881
- DOI: https://doi.org/10.1073/pnas.1216499110
- Primary Citation of Related Structures:
4G8O, 4G8R - PubMed Abstract:
Plasminogen activator inhibitor type-1 (PAI-1) is a member of the serine protease inhibitor (serpin) family. Excessive PAI-1 activity is associated with human disease, making it an attractive pharmaceutical target. However, like other serpins, PAI-1 has a labile structure, making it a difficult target for the development of small molecule inhibitors, and to date, there are no US Food and Drug Administration-approved small molecule inactivators of any serpins. Here we describe the mechanistic and structural characterization of a high affinity inactivator of PAI-1. This molecule binds to PAI-1 reversibly and acts through an allosteric mechanism that inhibits PAI-1 binding to proteases and to its cofactor vitronectin. The binding site is identified by X-ray crystallography and mutagenesis as a pocket at the interface of β-sheets B and C and α-helix H. A similar pocket is present on other serpins, suggesting that this site could be a common target in this structurally conserved protein family.
Organizational Affiliation:
Departments of Pathology and Internal Medicine, Division of Cardiovascular Medicine, University of Michigan Medical School, Ann Arbor, MI 48109.