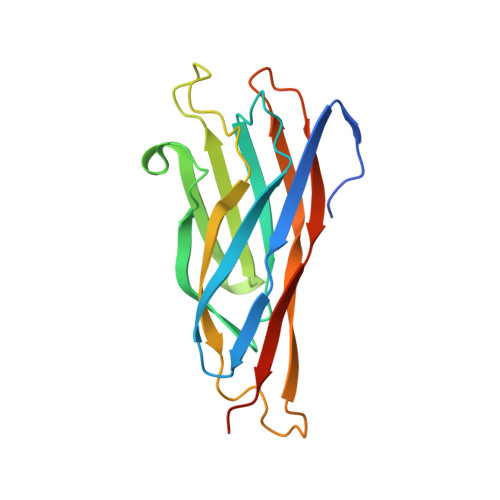
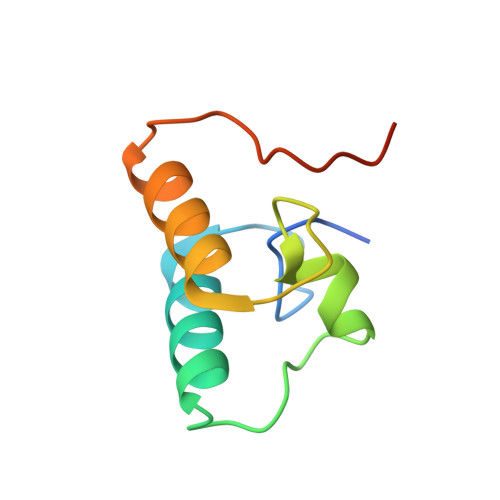
Novel Clostridium thermocellum Type I Cohesin-Dockerin Complexes Reveal a Single Binding Mode.
Bras, J.L., Alves, V.D., Carvalho, A.L., Najmudin, S., Prates, J.A., Ferreira, L.M., Bolam, D.N., Romao, M.J., Gilbert, H.J., Fontes, C.M.(2012) J Biol Chem 287: 44394-44405
- PubMed: 23118225
- DOI: https://doi.org/10.1074/jbc.M112.407700
- Primary Citation of Related Structures:
3UL4, 4DH2 - PubMed Abstract:
Protein-protein interactions play a pivotal role in a large number of biological processes exemplified by the assembly of the cellulosome. Integration of cellulosomal components occurs through the binding of type I cohesin modules located in a non-catalytic molecular scaffold to type I dockerin modules located at the C terminus of cellulosomal enzymes. The majority of type I dockerins display internal symmetry reflected by the presence of two essentially identical cohesin-binding surfaces. Here we report the crystal structures of two novel Clostridium thermocellum type I cohesin-dockerin complexes (CohOlpC-Doc124A and CohOlpA-Doc918). The data revealed that the two dockerins, Doc918 and Doc124A, are unusual because they lack the structural symmetry required to support a dual binding mode. Thus, in both cases, cohesin recognition is dominated by residues located at positions 11, 12, and 19 of one of the dockerin binding surfaces. The alternative binding mode is not possible (Doc918) or highly limited (Doc124A) because residues that assume the critical interacting positions, when dockerins are reoriented by 180°, make steric clashes with the cohesin. In common with a third dockerin (Doc258) that also presents a single binding mode, Doc124A directs the appended cellulase, Cel124A, to the surface of C. thermocellum and not to cellulosomes because it binds preferentially to type I cohesins located at the cell envelope. Although there are a few exceptions, such as Doc918 described here, these data suggest that there is considerable selective pressure for the evolution of a dual binding mode in type I dockerins that direct enzymes into cellulosomes.
Organizational Affiliation:
Centro de Investigação Interdisciplinar em Sanidade Animal, Faculdade de Medicina Veterinária, Universidade Técnica de Lisboa, Avenida da Universidade Técnica, 1300-477 Lisboa, Portugal.