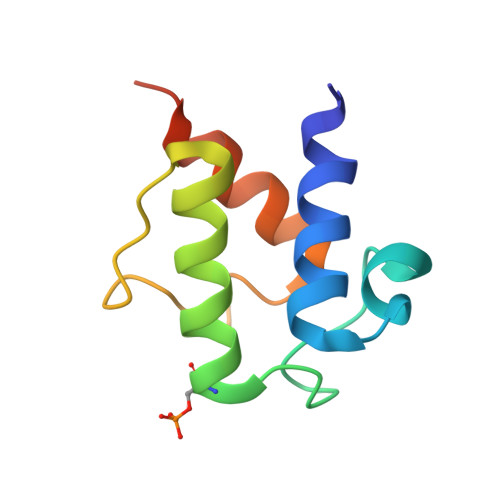
High-Resolution Structures of the D-Alanyl Carrier Protein (Dcp) Dltc from Bacillus Subtilis Reveal Equivalent Conformations of Apo- and Holo-Forms
Zimmermann, S., Pfennig, S., Neumann, P., Yonus, H., Weininger, U., Kovermann, M., Balbach, J., Stubbs, M.T.(2015) FEBS Lett 589: 2283
- PubMed: 26193422
- DOI: https://doi.org/10.1016/j.febslet.2015.07.008
- Primary Citation of Related Structures:
4BPF, 4BPG, 4BPH - PubMed Abstract:
D-Alanylation of lipoteichoic acids plays an important role in modulating the properties of Gram-positive bacteria cell walls. The D-alanyl carrier protein DltC from Bacillus subtilis has been solved in apo- and two cofactor-modified holo-forms, whereby the entire phosphopantetheine moiety is defined in one. The atomic resolution of the apo-structure allows delineation of alternative conformations within the hydrophobic core of the 78 residue four helix bundle. In contrast to previous reports for a peptidyl carrier protein from a non-ribosomal peptide synthetase, no obvious structural differences between apo- and holo-DltC forms are observed. Solution NMR spectroscopy confirms these findings and demonstrates in addition that the two forms exhibit similar backbone dynamics on the ps-ns and ms timescales.
Organizational Affiliation:
Institut für Biochemie und Biotechnologie, Martin-Luther Universität Halle-Wittenberg, Kurt-Mothes Strasse 3, D-06120 Halle/Saale, Germany.