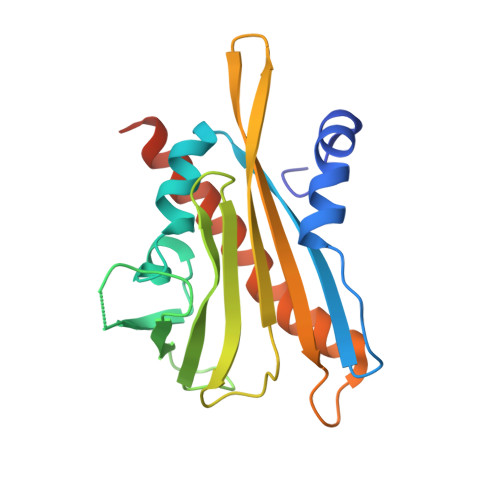
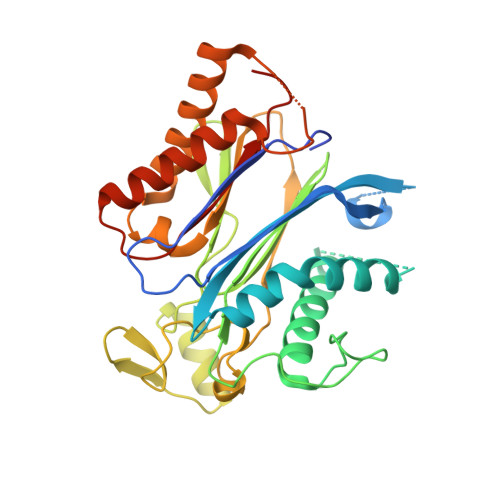
Modulation of Abscisic Acid Signaling in Vivo by an Engineered Receptor-Insensitive Protein Phosphatase Type 2C Allele.
Dupeux, F., Antoni, R., Betz, K., Santiago, J., Gonzalez-Guzman, M., Rodriguez, L., Rubio, S., Park, S.Y., Cutler, S.R., Rodriguez, P.L., Marquez, J.A.(2011) Plant Physiol 156: 106-116
- PubMed: 21357183
- DOI: https://doi.org/10.1104/pp.110.170894
- Primary Citation of Related Structures:
3QN1 - PubMed Abstract:
The plant hormone abscisic acid (ABA) plays a crucial role in the control of the stress response and the regulation of plant growth and development. ABA binding to PYRABACTIN RESISTANCE1 (PYR1)/PYR1-LIKE (PYL)/REGULATORY COMPONENTS OF ABA RECEPTORS intracellular receptors leads to inhibition of key negative regulators of ABA signaling, i.e. clade A protein phosphatases type 2C (PP2Cs) such as ABA-INSENSITIVE1 and HYPERSENSITIVE TO ABA1 (HAB1), causing the activation of the ABA signaling pathway. To gain further understanding on the mechanism of hormone perception, PP2C inhibition, and its implications for ABA signaling, we have performed a structural and functional analysis of the PYR1-ABA-HAB1 complex. Based on structural data, we generated a gain-of-function mutation in a critical residue of the phosphatase, hab1(W385A), which abolished ABA-dependent receptor-mediated PP2C inhibition without impairing basal PP2C activity. As a result, hab1(W385A) caused constitutive inactivation of the protein kinase OST1 even in the presence of ABA and PYR/PYL proteins, in contrast to the receptor-sensitive HAB1, and therefore hab1(W385A) qualifies as a hypermorphic mutation. Expression of hab1(W385A) in Arabidopsis (Arabidopsis thaliana) plants leads to a strong, dominant ABA insensitivity, which demonstrates that this conserved tryptophan residue can be targeted for the generation of dominant clade A PP2C alleles. Moreover, our data highlight the critical role of molecular interactions mediated by tryptophan-385 equivalent residues for clade A PP2C function in vivo and the mechanism of ABA perception and signaling.
Organizational Affiliation:
European Molecular Biology Laboratory, Grenoble Outstation and Unit of Virus Host-Cell Interactions, Université Joseph Fourier-EMBL-Centre National de la Recherche Scientifique, 38042 Grenoble cedex 9, France.