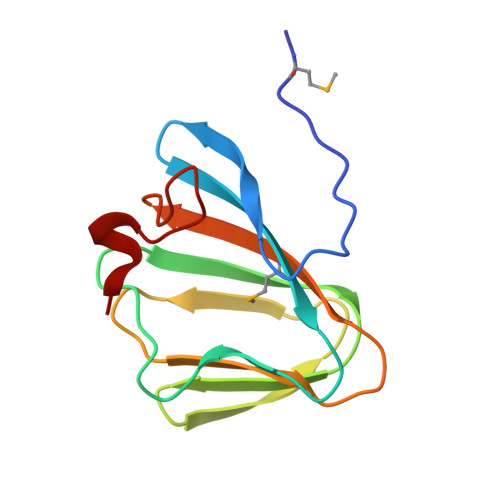
Conformational changes associated with the binding of zinc acetate at the putative active site of XcTcmJ, a cupin from Xanthomonas campestris pv. campestris.
Axelrod, H.L., Kozbial, P., McMullan, D., Krishna, S.S., Miller, M.D., Abdubek, P., Acosta, C., Astakhova, T., Carlton, D., Caruthers, J., Chiu, H.J., Clayton, T., Deller, M.C., Duan, L., Elias, Y., Feuerhelm, J., Grzechnik, S.K., Grant, J.C., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kumar, A., Marciano, D., Morse, A.T., Murphy, K.D., Nigoghossian, E., Okach, L., Oommachen, S., Paulsen, J., Reyes, R., Rife, C.L., Tien, H.J., Trout, C.V., van den Bedem, H., Weekes, D., White, A., Xu, Q., Zubieta, C., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1347-1353
- PubMed: 20944231
- DOI: https://doi.org/10.1107/S1744309109021988
- Primary Citation of Related Structures:
3H50 - PubMed Abstract:
In the plant pathogen Xanthomonas campestris pv. campestris, the product of the tcmJ gene, XcTcmJ, encodes a protein belonging to the RmlC family of cupins. XcTcmJ was crystallized in a monoclinic space group (C2) in the presence of zinc acetate and the structure was determined to 1.6 Å resolution. Previously, the apo structure has been reported in the absence of any bound metal ion [Chin et al. (2006), Proteins, 65, 1046-1050]. The most significant difference between the apo structure and the structure of XcTcmJ described here is a reorganization of the binding site for zinc acetate, which was most likely acquired from the crystallization solution. This site is located in the conserved metal ion-binding domain at the putative active site of XcTcmJ. In addition, an acetate was also bound within coordination distance of the zinc. In order to accommodate this binding, rearrangement of a conserved histidine ligand is required as well as several nearby residues within and around the putative active site. These observations indicate that binding of zinc serves a functional role in this cupin protein.
Organizational Affiliation:
Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.