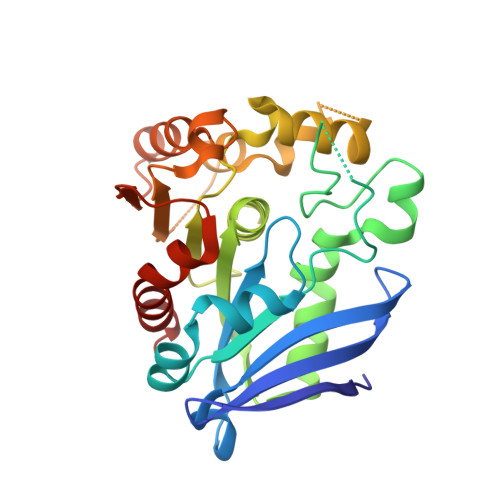
Crystal structure of human esterase D: a potential genetic marker of retinoblastoma
Wu, D., Li, Y., Song, G., Zhang, D., Shaw, N., Liu, Z.J.(2009) FASEB J 23: 1441-1446
- PubMed: 19126594
- DOI: https://doi.org/10.1096/fj.08-125286
- Primary Citation of Related Structures:
3FCX - PubMed Abstract:
Retinoblastoma (RB), a carcinoma of the retina, is caused by mutations in the long arm of chromosome 13, band 13q14. The esterase D (ESD) gene maps at a similar location as the RB gene locus and therefore serves as a potential marker for the prognosis of retinoblastoma. Because very little is known about the structure and function of ESD, we determined the 3-dimensional structure of the enzyme at 1.5 A resolution using X-ray crystallography. ESD shows a single domain with an alpha/beta-hydrolase fold. A number of insertions are observed in the canonical alpha/beta-hydrolase fold. The active site is located in a positively charged, shallow cleft on the surface lined by a number of aromatic residues. Superimposition studies helped identify the typical catalytic triad residues--Ser-153, His264, and Asp230--involved in catalysis. Mutagenesis of any of the catalytic triad residues to alanine abolished the enzyme activity. Backbone amides of Leu54 and Met150 are involved in the formation of the oxyanion hole. Interestingly, a M150A mutation increased the enzyme activity by 62%. The structure of human ESD determined in this study will aid the elucidation of the physiological role of the enzyme in the human body and will assist in the early diagnosis of retinoblastoma.
Organizational Affiliation:
National Laboratory of Biomacromolecules, Institute of Biophysics, 15 Datun Lu, Beijing 100101, China.