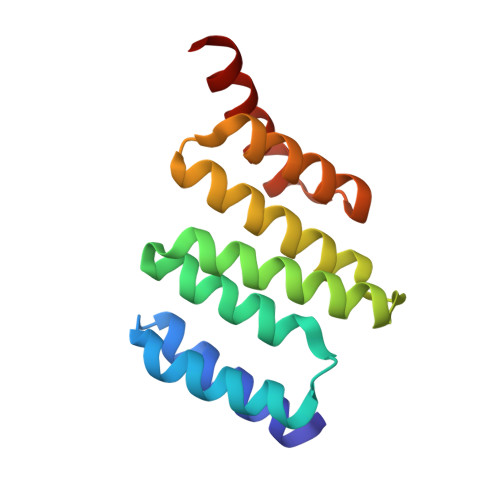
Electrostatic interactions of Hsp-organizing protein tetratricopeptide domains with Hsp70 and Hsp90: computational analysis and protein engineering.
Kajander, T., Sachs, J.N., Goldman, A., Regan, L.(2009) J Biol Chem 284: 25364-25374
- PubMed: 19586912
- DOI: https://doi.org/10.1074/jbc.M109.033894
- Primary Citation of Related Structures:
3ESK - PubMed Abstract:
The Hsp-organizing protein (HOP) binds to the C termini of the chaperones Hsp70 and Hsp90, thus bringing them together so that substrate proteins can be passed from Hsp70 to Hsp90. Because Hsp90 is essential for the correct folding and maturation of many oncogenic proteins, it has become a significant target for anti-cancer drug design. HOP binds to Hsp70 and Hsp90 via two independent tetratricopeptide (TPR) domains, TPR1 and TPR2A, respectively. We have analyzed ligand binding using Poisson-Boltzmann continuum electrostatic calculations, free energy perturbation, molecular dynamics simulations, and site-directed mutagenesis to delineate the contribution of different interactions to the affinity and specificity of the TPR-peptide interactions. We found that continuum electrostatic calculations could be used to guide protein design by removing unfavorable interactions to increase binding affinity, with an 80-fold increase in affinity for TPR2A. Contributions at buried charged residues, however, were better predicted by free energy perturbation calculations. We suggest using a combination of the two approaches for increasing the accuracy of results, with free energy perturbation calculations used only at selected buried residues of the ligand binding pocket. Finally we present the crystal structure of TPR2A in complex with its non-cognate Hsp70 ligand, which provides insight on the origins of specificity in TPR domain-peptide recognition.
Organizational Affiliation:
Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06511, USA. tommi.kajander@helsinki.fi