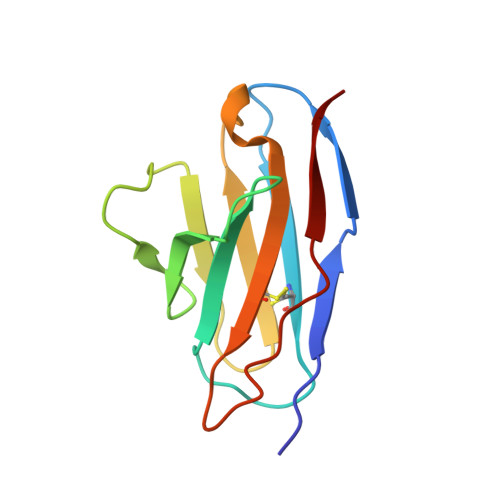
Structural alterations within native amyloidogenic immunoglobulin light chains.
Randles, E.G., Thompson, J.R., Martin, D.J., Ramirez-Alvarado, M.(2009) J Mol Biol 389: 199-210
- PubMed: 19361523
- DOI: https://doi.org/10.1016/j.jmb.2009.04.010
- Primary Citation of Related Structures:
3DVF, 3DVI - PubMed Abstract:
Amyloid diseases are characterized by the misfolding of a precursor protein that leads to amyloid fibril formation. Despite the fact that there are different precursors, some commonalities in the misfolding mechanism are thought to exist. In light chain amyloidosis (AL), the immunoglobulin light chain forms amyloid fibrils that deposit in the extracellular space of vital organs. AL proteins are thermodynamically destabilized compared to non-amyloidogenic proteins and some studies have linked this instability to increased fibril formation rates. Here we present the crystal structures of two highly homologous AL proteins, AL-12 and AL-103. This structural study shows that these proteins retain the canonical germ line dimer interface. We highlight important structural alterations in two loops flanking the dimer interface and correlate these results with the somatic mutations present in AL-12 and AL-103. We suggest that these alterations are informative structural features that are likely contributing to protein instability that leads to conformational changes involved in the initial events of amyloid formation.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, College of Medicine, Mayo Clinic, Rochester, MN 55905, USA.