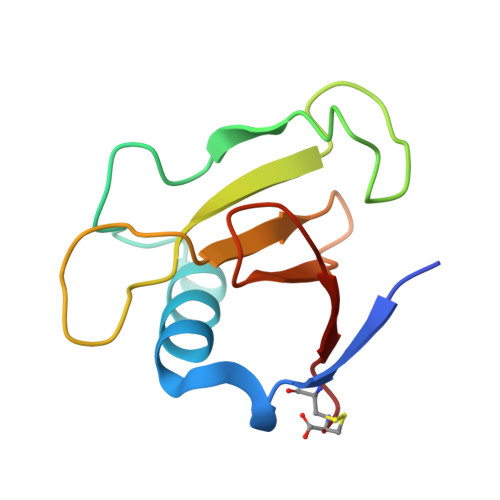
Structure of RNase Sa2 complexes with mononucleotides - new aspects of catalytic reaction and substrate recognition
Bauerova-Hlinkova, V., Dvorsky, R., Perecko, D., Povazanec, F., Sevcik, J.(2009) FEBS J 276: 4156-4168
- PubMed: 19558492
- DOI: https://doi.org/10.1111/j.1742-4658.2009.07125.x
- Primary Citation of Related Structures:
3D4A, 3D5G, 3D5I, 3DGY, 3DH2 - PubMed Abstract:
Although the mechanism of RNA cleavage by RNases has been studied for many years, there remain aspects that have not yet been fully clarified. We have solved the crystal structures of RNase Sa2 in the apo form and in complexes with mononucleotides. These structures provide more details about the mechanism of RNA cleavage by RNase Sa2. In addition to Glu56 and His86, which are the principal catalytic residues, an important role in the first reaction step of RNA cleavage also seems to be played by Arg67 and Arg71, which are located in the phosphate-binding site and form hydrogen bonds with the oxygens of the phosphate group of the mononucleotides. Their positive charge very likely causes polarization of the bonds between the oxygens and the phosphorus atom, leading to electron deficiency on the phosphorus atom and facilitating nucleophilic attack by O2' of the ribose on the phosphorus atom, leading to cyclophosphate formation. The negatively charged Glu56 is in position to attract the proton from O2' of the ribose. Extended molecular docking of mononucleotides, dinucleotides and trinucleotides into the active site of the enzyme allowed us to better understand the guanosine specificity of RNase Sa2 and to predict possible binding subsites for the downstream base and ribose of the second and third nucleotides.
Organizational Affiliation:
Institute of Molecular Biology, Slovak Academy of Sciences, Bratislava, Slovakia. vladena.hlinkova@savba.sk