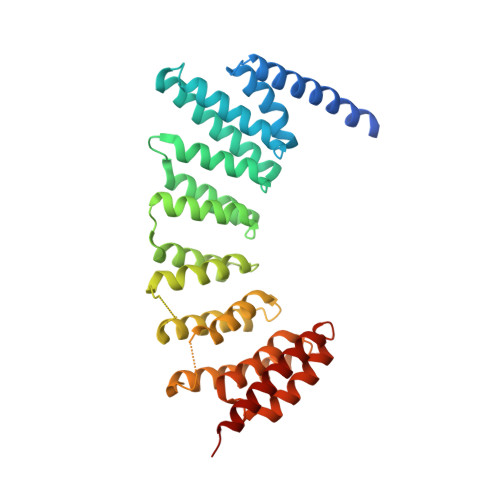
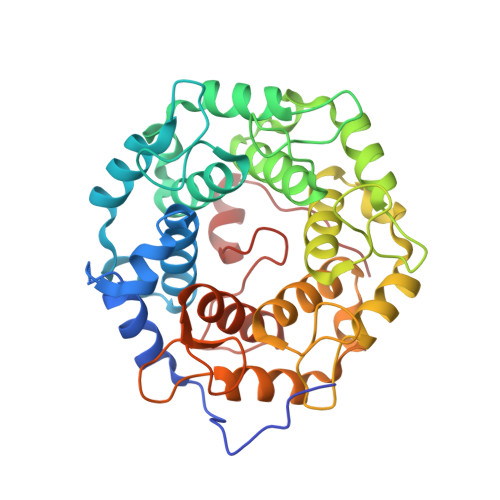
Development of selective RabGGTase inhibitors and crystal structure of a RabGGTase-inhibitor complex.
Guo, Z., Wu, Y.W., Tan, K.T., Bon, R.S., Guiu-Rozas, E., Delon, C., Nguyen, T.U., Wetzel, S., Arndt, S., Goody, R.S., Blankenfeldt, W., Alexandrov, K., Waldmann, H.(2008) Angew Chem Int Ed Engl 47: 3747-3750
- PubMed: 18399557
- DOI: https://doi.org/10.1002/anie.200705795
- Primary Citation of Related Structures:
3C72
Organizational Affiliation:
Max-Planck-Institut für molekulare Physiologie, Abt. Physikalische Biochemie, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.