NADP regulates the yeast GAL induction system.
Kumar, P.R., Yu, Y., Sternglanz, R., Johnston, S.A., Joshua-Tor, L.(2008) Science 319: 1090-1092
- PubMed: 18292341
- DOI: https://doi.org/10.1126/science.1151903
- Primary Citation of Related Structures:
3BTS, 3BTU, 3BTV - PubMed Abstract:
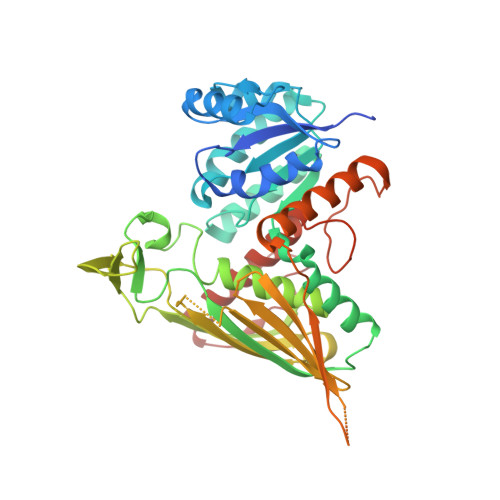
Transcriptional regulation of the galactose-metabolizing genes in Saccharomyces cerevisiae depends on three core proteins: Gal4p, the transcriptional activator that binds to upstream activating DNA sequences (UAS(GAL)); Gal80p, a repressor that binds to the carboxyl terminus of Gal4p and inhibits transcription; and Gal3p, a cytoplasmic transducer that, upon binding galactose and adenosine 5'-triphosphate, relieves Gal80p repression. The current model of induction relies on Gal3p sequestering Gal80p in the cytoplasm. However, the rapid induction of this system implies that there is a missing factor. Our structure of Gal80p in complex with a peptide from the carboxyl-terminal activation domain of Gal4p reveals the existence of a dinucleotide that mediates the interaction between the two. Biochemical and in vivo experiments suggests that nicotinamide adenine dinucleotide phosphate (NADP) plays a key role in the initial induction event.
Organizational Affiliation:
Cold Spring Harbor Laboratory, 1 Bungtown Road, Cold Spring Harbor, NY11724, USA.