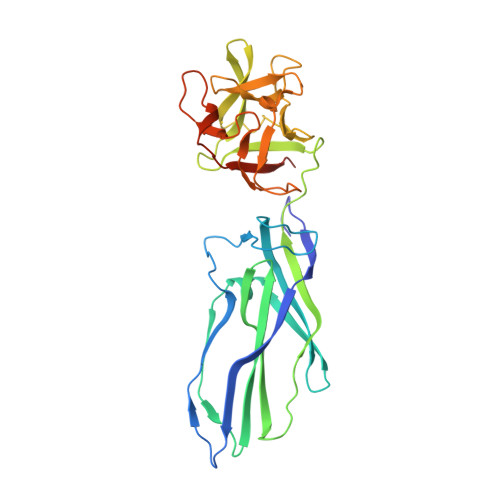
Structures of Lysenin Reveal a Shared Evolutionary Origin for Pore-Forming Proteins and its Mode of Sphingomyelin Recognition.
De Colibus, L., Sonnen, A.F.P., Morris, K.J., Siebert, C.A., Abrusci, P., Plitzko, J., Hodnik, V., Leippe, M., Volpi, E., Anderluh, G., Gilbert, R.J.C.(2012) Structure 20: 1498
- PubMed: 22819216
- DOI: https://doi.org/10.1016/j.str.2012.06.011
- Primary Citation of Related Structures:
3ZX7, 3ZXD, 3ZXG - PubMed Abstract:
Pore-forming proteins insert from solution into membranes to create lesions, undergoing a structural rearrangement often accompanied by oligomerization. Lysenin, a pore-forming toxin from the earthworm Eisenia fetida, specifically interacts with sphingomyelin (SM) and may confer innate immunity against parasites by attacking their membranes to form pores. SM has important roles in cell membranes and lysenin is a popular SM-labeling reagent. The structure of lysenin suggests common ancestry with other pore-forming proteins from a diverse set of eukaryotes and prokaryotes. The complex with SM shows the mode of its recognition by a protein in which both the phosphocholine headgroup and one acyl tail are specifically bound. Lipid interaction studies and assays using viable target cells confirm the functional reliance of lysenin on this form of SM recognition.
Organizational Affiliation:
Division of Structural Biology, University of Oxford, Roosevelt Drive, Oxford OX3 7BN, UK.