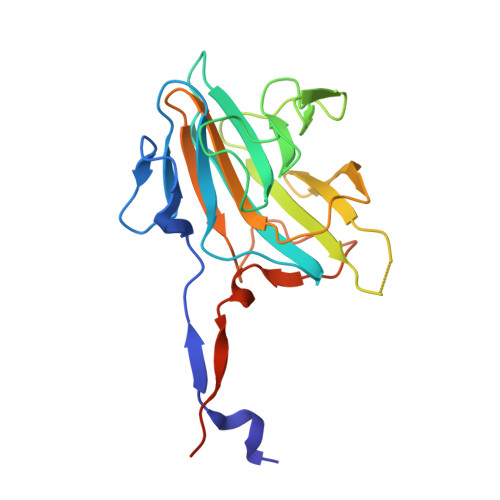
Structure of the SPRY domain of human Ash2L and its interactions with RbBP5 and DPY30.
Chen, Y., Cao, F., Wan, B., Dou, Y., Lei, M.(2012) Cell Res 22: 598-602
- PubMed: 22231628
- DOI: https://doi.org/10.1038/cr.2012.9
- Primary Citation of Related Structures:
3TOJ