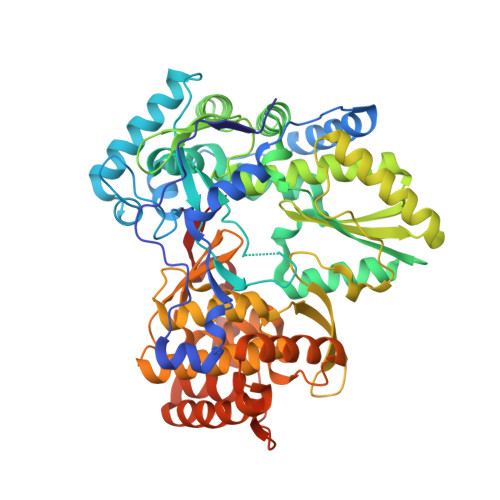
Investigation of the mode of binding of a novel series of N-benzyl-4-heteroaryl-1-(phenylsulfonyl)piperazine-2-carboxamides to the hepatitis C virus polymerase.
Gentles, R.G., Sheriff, S., Beno, B.R., Wan, C., Kish, K., Ding, M., Zheng, X., Chupak, L., Poss, M.A., Witmer, M.R., Morin, P., Wang, Y.K., Rigat, K., Lemm, J., Voss, S., Liu, M., Pelosi, L., Roberts, S.B., Gao, M., Kadow, J.F.(2011) Bioorg Med Chem Lett 21: 2212-2215
- PubMed: 21441029
- DOI: https://doi.org/10.1016/j.bmcl.2011.03.011
- Primary Citation of Related Structures:
3QGD, 3QGE, 3QGF, 3QGG, 3QGH, 3QGI - PubMed Abstract:
Structure based rationales for the activities of potent N-benzyl-4-heteroaryl-1-(phenylsulfonyl)piperazine-2-carboxamide inhibitors of the hepatitis C viral polymerase are described herein. These compounds bind to the hepatitis C virus non-structural protein 5B (NS5B), and co-crystal structures of select examples from this series with NS5B are reported. Comparison of co-crystal structures of a potent analog with both NS5B genotype 1a and genotype 1b provides a possible explanation for the genotype-selectivity observed with this compound class and suggests opportunities for the further optimization of the series.
Organizational Affiliation:
Bristol Myers Squibb, Chemical and Protein Technologies, Wallingford, CT 06492, United States. robert.gentles@bms.com