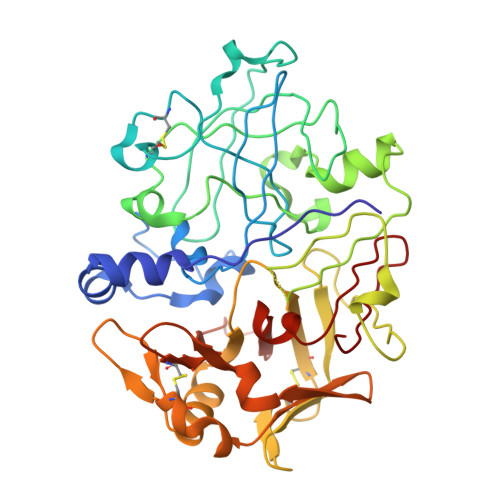
The high-resolution crystal structure of porcine pepsinogen.
Hartsuck, J.A., Koelsch, G., Remington, S.J.(1992) Proteins 13: 1-25
- PubMed: 1594574
- DOI: https://doi.org/10.1002/prot.340130102
- Primary Citation of Related Structures:
3PSG - PubMed Abstract:
The structure of porcine pepsinogen at pH 6.1 has been refined to an R-factor of 0.173 for data extending to 1.65 A. The final model contains 180 solvent molecules and lacks density for residues 157-161. The structure of this aspartic proteinase zymogen possesses many of the characteristics of pepsin, the mature enzyme. The secondary structure of the zymogen consists predominantly of beta-sheet, with an approximate 2-fold axis of symmetry. The activation peptide packs into the active site cleft, and the N-terminus (1P-9P) occupies the position of the mature N-terminus (1-9). Thus changes upon activation include excision of the activation peptide and proper relocation of the mature N-terminus. The activation peptide or residues of the displaced mature N-terminus make specific interactions with the substrate binding subsites. The active site of pepsinogen is intact; thus the lack of activity of pepsinogen is not due to a deformation of the active site. Nine ion pairs in pepsinogen may be important in the advent of activation and involve the activation peptide or regions of the mature N-terminus which are relocated in the mature enzyme. The activation peptide-pepsin junction, 44P-1, is characterized by high thermal parameters and weak density, indicating a flexible structure which would be accessible to cleavage. Pepsinogen is an appropriate model for the structures of other zymogens in the aspartic proteinase family.
Organizational Affiliation:
Protein Studies Program, Oklahoma Medical Research Foundation, University of Oklahoma Health Sciences Center, Oklahoma City 73104.