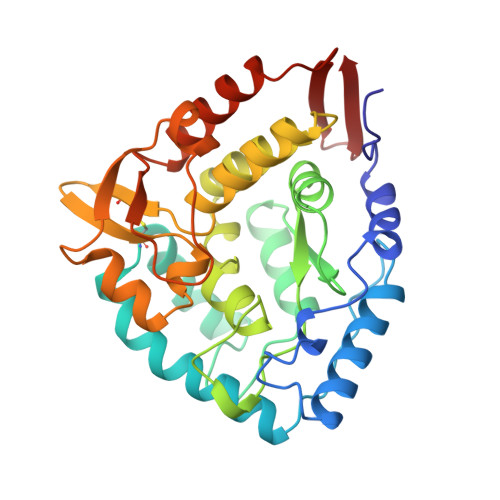
Crystallographic analysis of the human phenylalanine hydroxylase catalytic domain with bound catechol inhibitors at 2.0 A resolution.
Erlandsen, H., Flatmark, T., Stevens, R.C., Hough, E.(1998) Biochemistry 37: 15638-15646
- PubMed: 9843368
- DOI: https://doi.org/10.1021/bi9815290
- Primary Citation of Related Structures:
3PAH, 4PAH, 5PAH, 6PAH - PubMed Abstract:
The aromatic amino acid hydroxylases represent a superfamily of structurally and functionally closely related enzymes, one of those functions being reversible inhibition by catechol derivatives. Here we present the crystal structure of the dimeric catalytic domain (residues 117-424) of human phenylalanine hydroxylase (hPheOH), cocrystallized with various potent and well-known catechol inhibitors and refined at a resolution of 2.0 A. The catechols bind by bidentate coordination to each iron in both subunits of the dimer through the catechol hydroxyl groups, forming a blue-green colored ligand-to-metal charge-transfer complex. In addition, Glu330 and Tyr325 are identified as determinant residues in the recognition of the inhibitors. In particular, the interaction with Glu330 conforms to the structural explanation for the pH dependence of catecholamine binding to PheOH, with a pKa value of 5.1 (20 degreesC). The overall structure of the catechol-bound enzyme is very similar to that of the uncomplexed enzyme (rms difference of 0.2 A for the Calpha atoms). Most striking is the replacement of two iron-bound water molecules with catechol hydroxyl groups. This change is consistent with a change in the ligand field symmetry of the high-spin (S = 5/2) Fe(III) from a rhombic to a nearly axial ligand field symmetry as seen upon noradrenaline binding using EPR spectroscopy [Martinez, A., Andersson, K. K., Haavik, J., and Flatmark, T. (1991) Eur. J. Biochem. 198, 675-682]. Crystallographic comparison with the structurally related rat tyrosine hydroxylase binary complex with the oxidized cofactor 7,8-dihydrobiopterin revealed overlapping binding sites for the catechols and the cofactor, compatible with a competitive type of inhibition of the catechols versus BH4. The comparison demonstrates some structural differences at the active site as the potential basis for the different substrate specificity of the two enzymes.
Organizational Affiliation:
Protein Crystallography Group, Chemistry Department, University of Tromso, Norway.