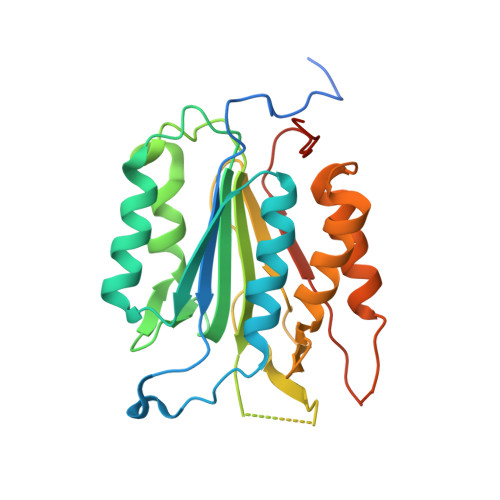
Crystal structures of human caspase 6 reveal a new mechanism for intramolecular cleavage self-activation
Wang, X.-J., Cao, Q., Liu, X., Wang, K.-T., Mi, W., Zhang, Y., Li, L.-F., Leblanc, A.C., Su, X.-D.(2010) EMBO Rep 11: 841-847
- PubMed: 20890311
- DOI: https://doi.org/10.1038/embor.2010.141
- Primary Citation of Related Structures:
3NR2, 3OD5 - PubMed Abstract:
Dimeric effectors caspase 3 and caspase 7 are activated by initiator caspase processing. In this study, we report the crystal structures of effector caspase 6 (CASP6) zymogen and N-Acetyl-Val-Glu-Ile-Asp-al-inhibited CASP6. Both of these forms of CASP6 have a dimeric structure, and in CASP6 zymogen the intersubunit cleavage site (190)TEVD(193) is well structured and inserts into the active site. This positions residue Asp 193 to be easily attacked by the catalytic residue Cys 163. We demonstrate biochemically that intramolecular cleavage at Asp 193 is a prerequisite for CASP6 self-activation and that this activation mechanism is dependent on the length of the L2 loop. Our results indicate that CASP6 can be activated and regulated through intramolecular self-cleavage.
Organizational Affiliation:
National Laboratory of Protein Engineering & Plant Genetic Engineering, Peking University, Haidian District, Beijing, People's Republic of China.