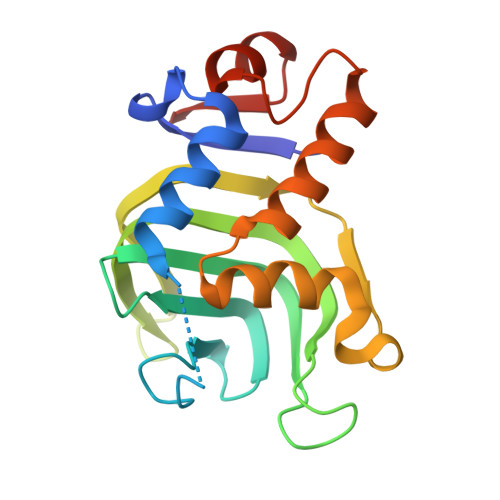
Structural, NMR Spectroscopic, and Computational Investigation of Hemin Loading in the Hemophore HasAp from Pseudomonas aeruginosa.
Jepkorir, G., Rodriguez, J.C., Rui, H., Im, W., Lovell, S., Battaile, K.P., Alontaga, A.Y., Yukl, E.T., Rivera, M.(2010) J Am Chem Soc 132: 9857-9872
- PubMed: 20572666
- DOI: https://doi.org/10.1021/ja103498z
- Primary Citation of Related Structures:
3MOK, 3MOL, 3MOM - PubMed Abstract:
When challenged by low-iron conditions several Gram-negative pathogens secrete a hemophore (HasA) to scavenge hemin from its host and deliver it to a receptor (HasR) on their outer membrane for internalization. Here we report results from studies aimed at probing the structural and dynamic processes at play in the loading of the apo-hemophore secreted by P. aeruginosa (apo-HasAp) with hemin. The structure of apo-HasAp shows a large conformational change in the loop harboring axial ligand His32 relative to the structure of holo-HasAp, whereas the loop bearing the other axial ligand, Tyr75, remains intact. To investigate the role played by the axial ligand-bearing loops in the process of hemin capture we investigated the H32A mutant, which was found to exist as a monomer in its apo-form and as a mixture of monomers and dimers in its holo-form. We obtained an X-ray structure of dimeric H32A holo-HasAp, which revealed that the two subunits are linked by cofacial interactions of two hemin molecules and that the conformation of the Ala32 loop in the dimer is identical to that exhibited by the His32 loop in wild type apo-HasAp. Additional data suggest that the conformation of the Ala32 loop in the dimer is mainly a consequence of dimerization. Hence, to investigate the effect of hemin loading on the topology of the His32 loop we also obtained the crystal structure of monomeric H32A holo-HasAp coordinated by imidazole (H32A-imidazole) and investigated the monomeric H32A HasAp and H32A-imidazole species in solution by NMR spectroscopy. The structure of H32A-imidazole revealed that the Ala32 loop attains a "closed" conformation nearly identical to that observed in wild type holo-HasAp, and the NMR investigations indicated that this conformation is maintained in solution. The NMR studies also highlighted conformational heterogeneity at the H32 loop hinges and in other key sections of the structure. Targeted molecular dynamics simulations allowed us to propose a possible path for the closing of the His32 loop upon hemin binding and identified molecular motions that are likely important in transmitting the presence of hemin in the Tyr75 loop to the His32 loop to initiate its closing. Importantly, residues implicated as undergoing motions in the computations are also observed as being dynamic by NMR. Taken together, these observations provide direct experimental evidence indicating that hemin loads onto the Tyr75 loop of apo-HasAp, which triggers the closing of the His32 loop.
Organizational Affiliation:
Department of Chemistry, University of Kansas, Multidisciplinary Research Building, 2030 Becker Drive, Room 220 E, Lawrence, Kansas 66047, USA.