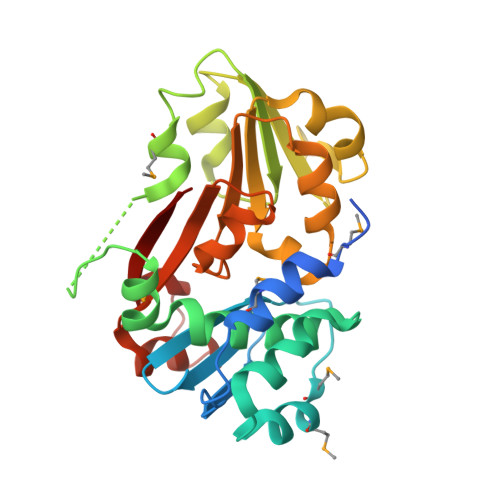
Structure of the first representative of Pfam family PF04016 (DUF364) reveals enolase and Rossmann-like folds that combine to form a unique active site with a possible role in heavy-metal chelation.
Miller, M.D., Aravind, L., Bakolitsa, C., Rife, C.L., Carlton, D., Abdubek, P., Astakhova, T., Axelrod, H.L., Chiu, H.J., Clayton, T., Deller, M.C., Duan, L., Feuerhelm, J., Grant, J.C., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Marciano, D., McMullan, D., Morse, A.T., Nigoghossian, E., Okach, L., Reyes, R., van den Bedem, H., Weekes, D., Xu, Q., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1167-1173
- PubMed: 20944207
- DOI: https://doi.org/10.1107/S1744309110007517
- Primary Citation of Related Structures:
3L5O - PubMed Abstract:
The crystal structure of Dhaf4260 from Desulfitobacterium hafniense DCB-2 was determined by single-wavelength anomalous diffraction (SAD) to a resolution of 2.01 Å using the semi-automated high-throughput pipeline of the Joint Center for Structural Genomics (JCSG) as part of the NIGMS Protein Structure Initiative (PSI). This protein structure is the first representative of the PF04016 (DUF364) Pfam family and reveals a novel combination of two well known domains (an enolase N-terminal-like fold followed by a Rossmann-like domain). Structural and bioinformatic analyses reveal partial similarities to Rossmann-like methyltransferases, with residues from the enolase-like fold combining to form a unique active site that is likely to be involved in the condensation or hydrolysis of molecules implicated in the synthesis of flavins, pterins or other siderophores. The genome context of Dhaf4260 and homologs additionally supports a role in heavy-metal chelation.
Organizational Affiliation:
Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.