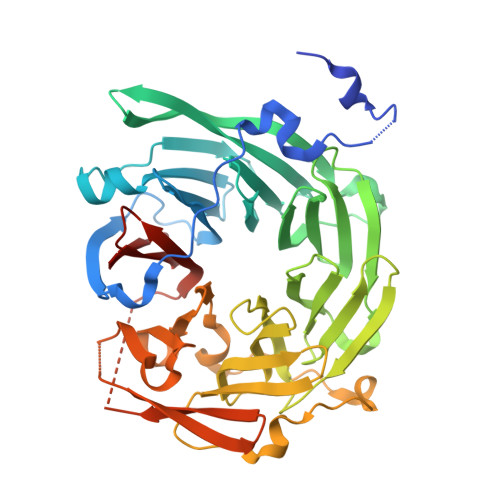
Structure and function of Vps15 in the endosomal G protein signaling pathway.
Heenan, E.J., Vanhooke, J.L., Temple, B.R., Betts, L., Sondek, J.E., Dohlman, H.G.(2009) Biochemistry 48: 6390-6401
- PubMed: 19445518
- DOI: https://doi.org/10.1021/bi900621w
- Primary Citation of Related Structures:
3GRE - PubMed Abstract:
G protein-coupled receptors mediate cellular responses to a wide variety of stimuli, including taste, light, and neurotransmitters. In the yeast Saccharomyces cerevisiae, activation of the pheromone pathway triggers events leading to mating. The view had long been held that the G protein-mediated signal occurs principally at the plasma membrane. Recently, it has been shown that the G protein alpha subunit Gpa1 can promote signaling at endosomes and requires two components of the sole phosphatidylinositol-3-kinase in yeast, Vps15 and Vps34. Vps15 contains multiple WD repeats and also binds to Gpa1 preferentially in the GDP-bound state; these observations led us to hypothesize that Vps15 may function as a G protein beta subunit at the endosome. Here we show an X-ray crystal structure of the Vps15 WD domain that reveals a seven-bladed propeller resembling that of typical Gbeta subunits. We show further that the WD domain is sufficient to bind Gpa1 as well as to Atg14, a potential Ggamma protein that exists in a complex with Vps15. The Vps15 kinase domain together with the intermediate domain (linking the kinase and WD domains) also contributes to Gpa1 binding and is necessary for Vps15 to sustain G protein signaling. These findings reveal that the Vps15 Gbeta-like domain serves as a scaffold to assemble Gpa1 and Atg14, whereas the kinase and intermediate domains are required for proper signaling at the endosome.
Organizational Affiliation:
Department of Biochemistry and Biophysics, University of North Carolina, Chapel Hill, North Carolina 27599-7260, USA.