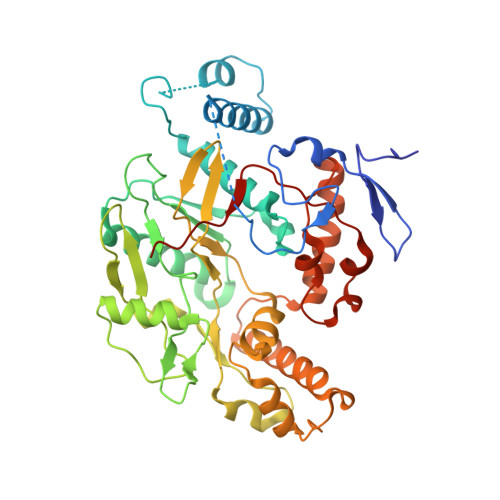
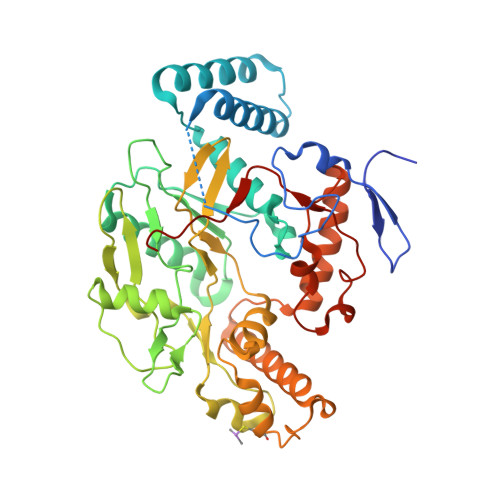
Anchored plasticity opens doors for selective inhibitor design in nitric oxide synthase.
Garcin, E.D., Arvai, A.S., Rosenfeld, R.J., Kroeger, M.D., Crane, B.R., Andersson, G., Andrews, G., Hamley, P.J., Mallinder, P.R., Nicholls, D.J., St-Gallay, S.A., Tinker, A.C., Gensmantel, N.P., Mete, A., Cheshire, D.R., Connolly, S., Stuehr, D.J., Aberg, A., Wallace, A.V., Tainer, J.A., Getzoff, E.D.(2008) Nat Chem Biol 4: 700-707
- PubMed: 18849972
- DOI: https://doi.org/10.1038/nchembio.115
- Primary Citation of Related Structures:
3E65, 3E67, 3E6N, 3E6O, 3E6T, 3E7G, 3E7I, 3E7M, 3E7S, 3E7T, 3EAH, 3EAI, 3EBD, 3EBF, 3EJ8 - PubMed Abstract:
Nitric oxide synthase (NOS) enzymes synthesize nitric oxide, a signal for vasodilatation and neurotransmission at low concentrations and a defensive cytotoxin at higher concentrations. The high active site conservation among all three NOS isozymes hinders the design of selective NOS inhibitors to treat inflammation, arthritis, stroke, septic shock and cancer. Our crystal structures and mutagenesis results identified an isozyme-specific induced-fit binding mode linking a cascade of conformational changes to a new specificity pocket. Plasticity of an isozyme-specific triad of distant second- and third-shell residues modulates conformational changes of invariant first-shell residues to determine inhibitor selectivity. To design potent and selective NOS inhibitors, we developed the anchored plasticity approach: anchor an inhibitor core in a conserved binding pocket, then extend rigid bulky substituents toward remote specificity pockets, which become accessible upon conformational changes of flexible residues. This approach exemplifies general principles for the design of selective enzyme inhibitors that overcome strong active site conservation.
Organizational Affiliation:
The Scripps Research Institute, Department of Molecular Biology and Skaggs Institute for Chemical Biology, 10550 North Torrey Pines Road, MB4, La Jolla, California 92037, USA. egarcin@umbc.edu