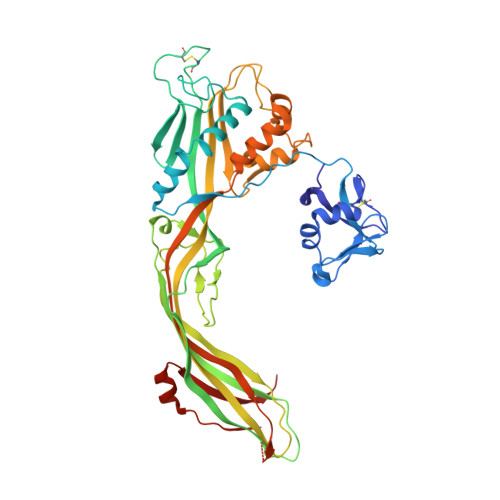
Molecular assembly of the aerolysin pore reveals a swirling membrane-insertion mechanism.
Degiacomi, M.T., Iacovache, I., Pernot, L., Chami, M., Kudryashev, M., Stahlberg, H., van der Goot, F.G., Dal Peraro, M.(2013) Nat Chem Biol 9: 623-629
- PubMed: 23912165
- DOI: https://doi.org/10.1038/nchembio.1312
- Primary Citation of Related Structures:
3C0M, 3C0N - PubMed Abstract:
Aerolysin is the founding member of a superfamily of β-pore-forming toxins whose pore structure is unknown. We have combined X-ray crystallography, cryo-EM, molecular dynamics and computational modeling to determine the structures of aerolysin mutants in their monomeric and heptameric forms, trapped at various stages of the pore formation process. A dynamic modeling approach based on swarm intelligence was applied, whereby the intrinsic flexibility of aerolysin extracted from new X-ray structures was used to fully exploit the cryo-EM spatial restraints. Using this integrated strategy, we obtained a radically new arrangement of the prepore conformation and a near-atomistic structure of the aerolysin pore, which is fully consistent with all of the biochemical data available so far. Upon transition from the prepore to pore, the aerolysin heptamer shows a unique concerted swirling movement, accompanied by a vertical collapse of the complex, ultimately leading to the insertion of a transmembrane β-barrel.
Organizational Affiliation:
1] Institute of Bioengineering, School of Life Sciences, Ecole Polytechnique Fédérale de Lausanne, Lausanne, Switzerland. [2] [3].