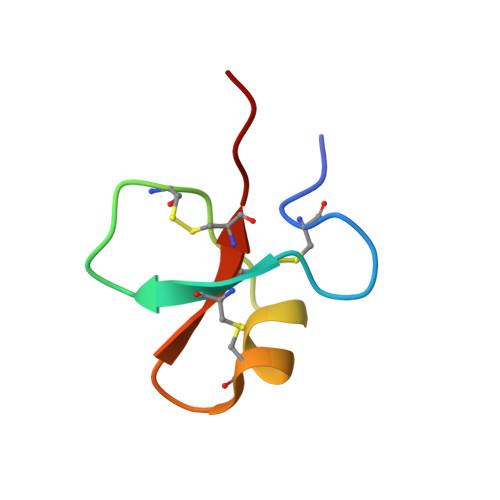
The NMR structures of the major intermediates of the two-domain tick carboxypeptidase inhibitor reveal symmetry in its folding and unfolding pathways.
Arolas, J.L., Pantoja-Uceda, D., Ventura, S., Blanco, F.J., Aviles, F.X.(2008) J Biol Chem 283: 27110-27120
- PubMed: 18640980
- DOI: https://doi.org/10.1074/jbc.M803978200
- Primary Citation of Related Structures:
2K2X, 2K2Y, 2K2Z - PubMed Abstract:
There is a lack of experimental structural information about folding intermediates of multidomain proteins. Tick carboxypeptidase inhibitor (TCI) is a small, disulfide-rich protein consisting of two domains that fold and unfold autonomously through the formation of two major intermediates, IIIa and IIIb. Each intermediate contains three native disulfide bonds in one domain and six free cysteines in the other domain. Here we have determined the NMR structures of these two intermediates trapped and isolated at acidic pH in which they are stable and compared their structures with that of the native protein analyzed under the same conditions. Both IIIa and IIIb were found to contain a folded region that corresponds to the N- and C-terminal domains of TCI, respectively, with structures very similar to the corresponding regions of the native protein. The remainder of the polypeptide chains of the intermediates was shown to be unfolded in a random coil conformation. Solvent exchange measurements further indicated that the two protein domains are not completely independent, but affect each other in terms of dynamics and stability, in agreement with reported inhibitory activity data. The derived results provide structural evidence for symmetric TCI folding and unfolding mechanisms that converge in IIIa and IIIb and reveal the structural basis that accounts for the strong and simultaneous accumulation of both intermediates. Altogether, this work has important implications for a better understanding of the folding mechanisms of multidomain, disulfide-rich proteins.
Organizational Affiliation:
Institut de Biotecnologia i Biomedicina and Departament de Bioquímica i Biologia Molecular, Universitat Autònoma de Barcelona, 08193 Bellaterra, Spain.