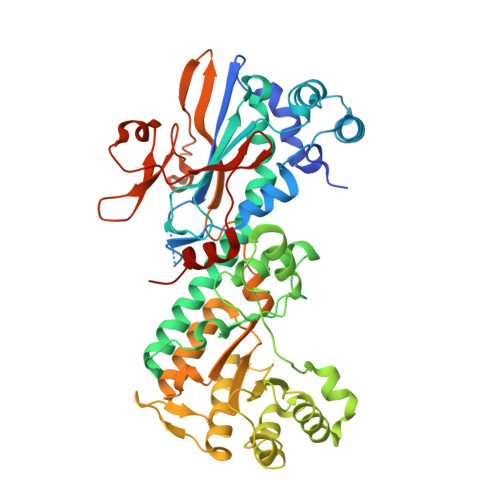
Structure and reaction mechanism of human nicotinamide phosphoribosyltransferase
Takahashi, R., Nakamura, S., Nakazawa, T., Minoura, K., Yoshida, T., Nishi, Y., Kobayashi, Y., Ohkubo, T.(2010) J Biochem 147: 95-107
- PubMed: 19819904
- DOI: https://doi.org/10.1093/jb/mvp152
- Primary Citation of Related Structures:
2E5B, 2E5C, 2E5D - PubMed Abstract:
Nicotinamide (NM) phosphoribosyltransferase (NMPRTase) catalyzes the reaction of NM and 5'-phosphoribosyl-1'-pyrophosphate (PRPP) to form NM mononucleotide (NMN) and pyrophosphate (PPi) in the pathway of NAD-biosynthesis. Monitoring the (1)H and (31)P NMR spectra of the reaction mixture, we found that this reaction is reversible as dictated by the equilibrium constant K = [NMN][PPi]/([NM][PRPP]) = 0.14, which agreed well with the ratio of second-order rate constants for forward and backward reactions, K = 0.16. The crystal structures of this enzyme in the free form and bound to NM and PRPP at the resolution of 2.0-2.2 A were essentially identical to that of the complex with NMN, except for some variations that could facilitate the substitution reaction by fixing the nucleophile and the leaving group for the requisite inversion of configuration at the C1' carbon of the ribose ring. In the active site near the C1' atom of the bound PRPP or NMN, there was neither negatively charged group nor waterproof environment necessary to support the feasibility of a ribo-oxocarbocation intermediate inherent in the S(N)1 mechanism. The structures and catalytic mechanism thus revealed are also discussed in connection with the multiple biological functions of NMPRTase.
Organizational Affiliation:
Graduate School of Pharmaceutical Sciences, Osaka University, Suita, Osaka, Japan.