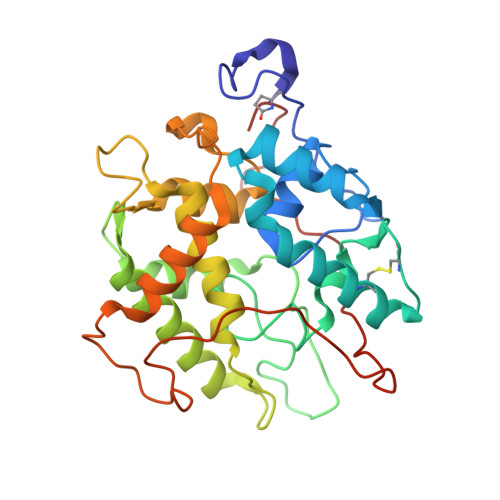
Crystal Structures of Chloroperoxidase with its Bound Substrates and Complexed with Formate, Acetate, and Nitrate.
Kuhnel, K., Blankenfeldt, W., Terner, J., Schlichting, I.(2006) J Biol Chem 281: 23990
- PubMed: 16790441
- DOI: https://doi.org/10.1074/jbc.M603166200
- Primary Citation of Related Structures:
2CIV, 2CIW, 2CIX, 2CIY, 2CIZ, 2CJ0, 2CJ1, 2CJ2 - PubMed Abstract:
Chloroperoxidase (CPO) is a heme-thiolate enzyme that catalyzes hydrogen peroxide-dependent halogenation reactions. Structural data on substrate binding have not been available so far. CPO was therefore crystallized in the presence of iodide or bromide. One halide binding site was identified at the surface near a narrow channel that connects the surface with the heme. Two other halide binding sites were identified within and at the other end of this channel. Together, these sites suggest a pathway for access of halide anions to the active site. The structure of CPO complexed with its natural substrate cyclopentanedione was determined at a resolution of 1.8 A. This is the first example of a CPO structure with a bound organic substrate. In addition, structures of CPO bound with nitrate, acetate, and formate and of a ternary complex with dimethylsulfoxide (Me2SO) and cyanide were determined. These structures have implications for the mechanism of compound I formation. Before binding to the heme, the incoming hydrogen peroxide first interacts with Glu-183. The deprotonated Glu-183 abstracts a proton from hydrogen peroxide. The hydroperoxo-anion then binds at the heme, yielding compound 0. Glu-183 protonates the distal oxygen of compound 0, water is released, and compound I is formed.
Organizational Affiliation:
Abteilung Biomolekulare Mechanismen, Max-Planck-Institut für Medizinische Forschung, Jahnstrasse 29, 69120 Heidelberg, Germany.