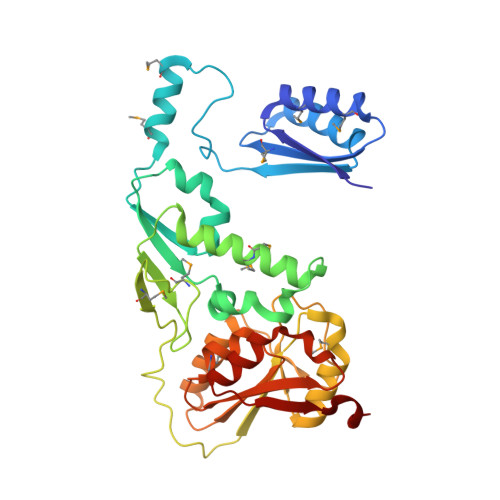
Structure and Mechanism of the Lipooligosaccharide Sialyltransferase from Neisseria Meningitidis
Lin, L.Y.C., Rakic, B., Chiu, C.P.C., Lameignere, E., Wakarchuk, W.W., Withers, S.G., Strynadka, N.C.J.(2011) J Biol Chem 286: 37237
- PubMed: 21880735
- DOI: https://doi.org/10.1074/jbc.M111.249920
- Primary Citation of Related Structures:
2YK4, 2YK5, 2YK6, 2YK7 - PubMed Abstract:
The first x-ray crystallographic structure of a CAZY family-52 glycosyltransferase, that of the membrane associated α2,3/α2,6 lipooligosaccharide sialyltransferase from Neisseria meningitidis serotype L1 (NST), has been solved to 1.95 Å resolution. The structure of NST adopts a GT-B-fold common with other glycosyltransferase (GT) families but exhibits a novel domain swap of the N-terminal 130 residues to create a functional homodimeric form not observed in any other class to date. The domain swap is mediated at the structural level by a loop-helix-loop extension between residues Leu-108 and Met-130 (we term the swapping module) and a unique lipid-binding domain. NST catalyzes the creation of α2,3- or 2,6-linked oligosaccharide products from a CMP-sialic acid (Neu5Ac) donor and galactosyl-containing acceptor sugars. Our structures of NST bound to the non-hydrolyzable substrate analog CMP-3F((axial))-Neu5Ac show that the swapping module from one monomer of NST mediates the binding of the donor sugar in a composite active site formed at the dimeric interface. Kinetic analysis of designed point mutations observed in the CMP-3F((axial))-Neu5Ac binding site suggests potential roles of a requisite general base (Asp-258) and general acid (His-280) in the NST catalytic mechanism. A long hydrophobic tunnel adjacent to the dimer interface in each of the two monomers contains electron density for two extended linear molecules that likely belong to either the two fatty acyl chains of a diglyceride lipid or the two polyethylene glycol groups of the detergent Triton X-100. In this work, Triton X-100 maintains the activity and increases the solubility of NST during purification and is critical to the formation of ordered crystals. Together, the mechanistic implications of the NST structure provide insight into lipooligosaccharide sialylation with respect to the association of substrates and the essential membrane-anchored nature of NST on the bacterial surface.
Organizational Affiliation:
Department of Biochemistry and Molecular Biology, Centre for Blood Research University of British Columbia, Vancouver, British Columbia V6T 1Z3, Canada.