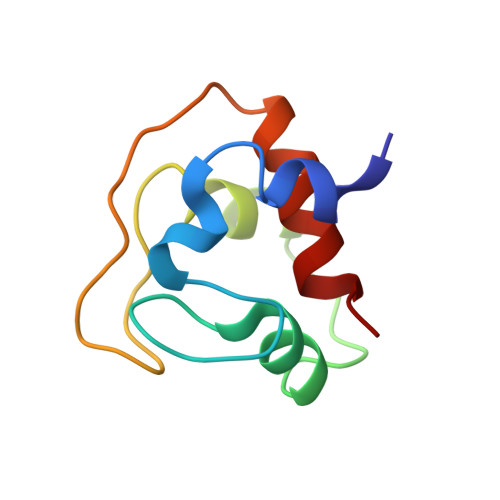
Solution structure of Fe(II) cytochrome c551 from Pseudomonas aeruginosa as determined by two-dimensional 1H NMR.
Detlefsen, D.J., Thanabal, V., Pecoraro, V.L., Wagner, G.(1991) Biochemistry 30: 9040-9046
- PubMed: 1654086
- DOI: https://doi.org/10.1021/bi00101a019
- Primary Citation of Related Structures:
2PAC - PubMed Abstract:
The solution structure of Fe(II) cytochrome c551 from Pseudomonas aeruginosa based on 2D 1H NMR data is reported. Two sets of structure calculations were completed with a combination of simulated annealing and distance geometry calculations: one set of 20 structures included the heme-peptide covalent linkages, and one set of 10 structures excluded them. The main-chain atoms were well constrained within the two structural ensembles (1.30 and 1.35 A average RMSD, respectively) except for two regions spanning residues 30-40 and 60-70. The results were essentially the same when global fold comparisons were made between the ensembles with an average RMSD of 1.33 A. In total, 556 constraints were used, including 479 NOEs, 53 volume constraints, and 24 other distances. This report represents the first solution structure determination of a heme protein by 2D 1H NMR and should provide a basis for the application of these techniques to other proteins containing large prosthetic groups or cofactors.
Organizational Affiliation:
Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts 02115.