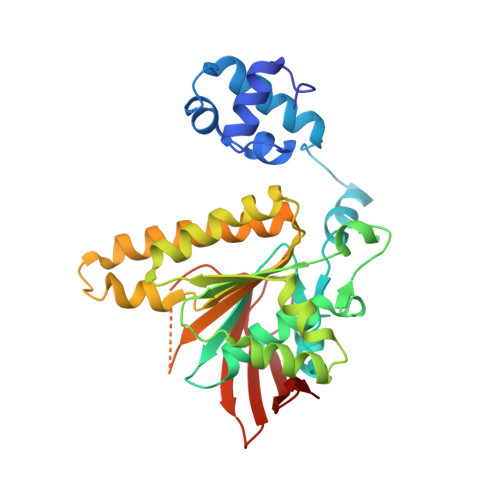
Crystal Structure of Methanococcus Voltae Rada in Complex with Adp: hydrolysis-induced conformational change
Qian, X., Wu, Y., He, Y., Moya, I.A., Luo, Y.(2005) Biochemistry 44: 13753-13761
- PubMed: 16229465
- DOI: https://doi.org/10.1021/bi051222i
- Primary Citation of Related Structures:
2FPK, 2FPL, 2FPM - PubMed Abstract:
Members of a superfamily of RecA-like recombinases facilitate a central strand exchange reaction in the DNA repair process. Archaeal RadA and Rad51 and eukaryal Rad51 and meiosis-specific DMC1 form a closely related group of recombinases distinct from bacterial RecA. Nevertheless, all such recombinases share a conserved core domain which carries the ATPase site and putative DNA-binding sites. Here we present the crystal structure of an archaeal RadA from Methanococcus voltae (MvRadA) in complex with ADP and Mg2+ at 2.1 A resolution. The crystallized RadA-ADP filament has an extended helical pitch similar to those of previously determined structures in the presence of nonhydrolyzable ATP analogue AMP-PNP. Structural comparison reveals two recurrent conformations with an extensive allosteric effect spanning the ATPase site and the putative DNA-binding L2 region. Varied conformations of the L2 region also imply a dynamic nature of recombinase-bound DNA.
Organizational Affiliation:
Department of Biochemistry, University of Saskatchewan, A3 Health Sciences Building, 107 Wiggins Road, Saskatoon, Saskatchewan, Canada S7N 5E5.