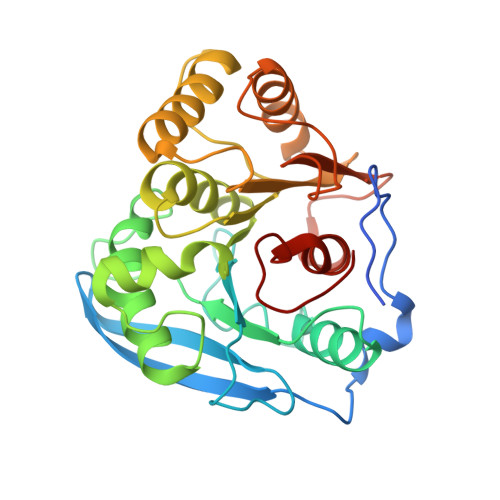
Molecular Determinants of Substrate Specificity in the Feruloyl Esterase Module of Xylanase 10B from Clostridium Thermocellum
Tarbouriech, N., Prates, J.A., Fontes, C., Davies, G.J.(2005) Acta Crystallogr D Biol Crystallogr 61: 194
- PubMed: 15681871
- DOI: https://doi.org/10.1107/S0907444904029695
- Primary Citation of Related Structures:
1WB4, 1WB5, 1WB6 - PubMed Abstract:
Feruloyl esterases play a key role in the degradation of the intricate structure of the plant cell wall by hydrolysing the ferulate ester groups involved in the cross-linking between hemicelluloses and between hemicellulose and lignin. The structure of the feruloyl esterase module of Clostridium thermocellum cellulosomal xylanase 10B has been reported previously. It displays the alpha/beta hydrolase fold with a classical Ser-His-Asp catalytic triad. Here, the structures of a Ser-Ala mutant of this feruloyl esterase in complexes with methyl syringate, methyl sinapinate and methyl vanillate are described. Substrate binding is accompanied by subtle conformational changes at amino acids Trp982, Met955, Asn1023 and Ile1019 in the ligand-binding cavity. The structural determinants, particularly the m-methoxy substituent, governing the substrate specificity of Xyn10B feruloyl esterase are rationalized.
Organizational Affiliation:
Structural Biology Laboratory, Department of Chemistry, University of York, Heslington, York YO10 5YW, England.