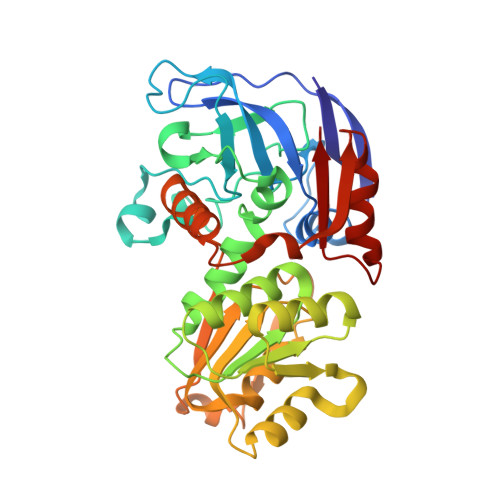
Structural basis for the enhanced thermal stability of alcohol dehydrogenase mutants from the mesophilic bacterium Clostridium beijerinckii: contribution of salt bridging
Bogin, O., Levin, I., Hacham, Y., Tel-Or, S., Peretz, M., Frolow, F., Burstein, Y.(2002) Protein Sci 11: 2561-2574
- PubMed: 12381840
- DOI: https://doi.org/10.1110/ps.0222102
- Primary Citation of Related Structures:
1JQB - PubMed Abstract:
Previous research in our laboratory comparing the three-dimensional structural elements of two highly homologous alcohol dehydrogenases, one from the mesophile Clostridium beijerinckii (CbADH) and the other from the extreme thermophile Thermoanaerobacter brockii (TbADH), suggested that in the thermophilic enzyme, an extra intrasubunit ion pair (Glu224-Lys254) and a short ion-pair network (Lys257-Asp237-Arg304-Glu165) at the intersubunit interface might contribute to the extreme thermal stability of TbADH. In the present study, we used site-directed mutagenesis to replace these structurally strategic residues in CbADH with the corresponding amino acids from TbADH, and we determined the effect of such replacements on the thermal stability of CbADH. Mutations in the intrasubunit ion pair region increased thermostability in the single mutant S254K- and in the double mutant V224E/S254K-CbADH, but not in the single mutant V224E-CbADH. Both single amino acid replacements, M304R- and Q165E-CbADH, in the region of the intersubunit ion pair network augmented thermal stability, with an additive effect in the double mutant M304R/Q165E-CbADH. To investigate the precise mechanism by which such mutations alter the molecular structure of CbADH to achieve enhanced thermostability, we constructed a quadruple mutant V224E/S254K/Q165E/M304R-CbADH and solved its three-dimensional structure. The overall results indicate that the amino acid substitutions in CbADH mutants with enhanced thermal stability reinforce the quaternary structure of the enzyme by formation of an extended network of intersubunit ion pairs and salt bridges, mediated by water molecules, and by forming a new intrasubunit salt bridge.
Organizational Affiliation:
Department of Organic Chemistry, The Weizmann Institute of Science, Rehovot 76100, Israel.