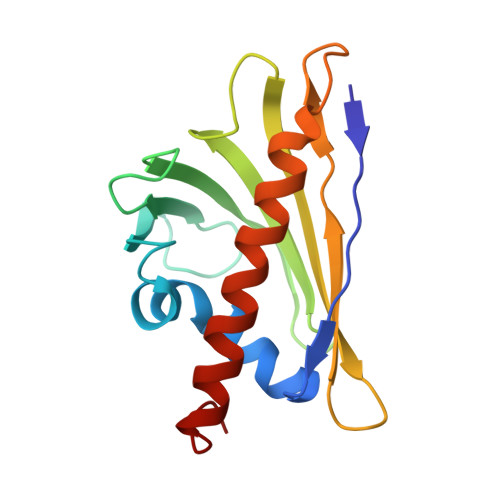
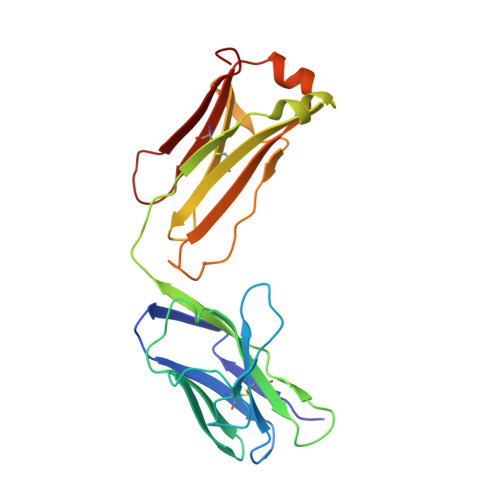
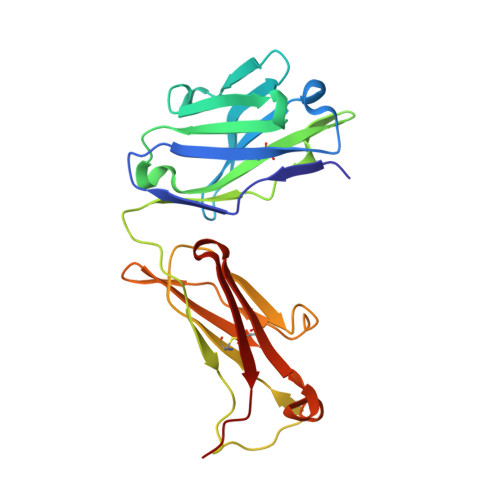
Dominant epitopes and allergic cross-reactivity: complex formation between a Fab fragment of a monoclonal murine IgG antibody and the major allergen from birch pollen Bet v 1.
Mirza, O., Henriksen, A., Ipsen, H., Larsen, J.N., Wissenbach, M., Spangfort, M.D., Gajhede, M.(2000) J Immunol 165: 331-338
- PubMed: 10861069
- DOI: https://doi.org/10.4049/jimmunol.165.1.331
- Primary Citation of Related Structures:
1FSK - PubMed Abstract:
The symptoms characteristic of allergic hypersensitivity are caused by the release of mediators, i.e., histamine, from effector cells such as basophils and mast cells. Allergens with more than one B cell epitope cross-link IgE Abs bound to high affinity FcepsilonRI receptors on mast cell surfaces leading to aggregation and subsequent mediator release. Thus, allergen-Ab complexes play a crucial role in the cascade leading to the allergic response. We here report the structure of a 1:1 complex between the major birch pollen allergen Bet v 1 and the Fab fragment from a murine monoclonal IgG1 Ab, BV16, that has been solved to 2.9 A resolution by x-ray diffraction. The mAb is shown to inhibit the binding of allergic patients' IgE to Bet v 1, and the allergen-IgG complex may therefore serve as a model for the study of allergen-IgE interactions relevant in allergy. The size of the BV16 epitope is 931 A2 as defined by the Bet v 1 Ab interaction surface. Molecular interactions predicted to occur in the interface are likewise in agreement with earlier observations on Ag-Ab complexes. The epitope is formed by amino acids that are conserved among major allergens from related species within the Fagales order. In combination with a surprisingly high inhibitory capacity of BV16 with respect to allergic patients' serum IgE binding to Bet v 1, these observations provide experimental support for the proposal of dominant IgE epitopes located in the conserved surface areas. This model will facilitate the development of new and safer vaccines for allergen immunotherapy in the form of mutated allergens.
Organizational Affiliation:
Protein Structure Group, Department of Chemistry, University of Copenhagen, Denmark.