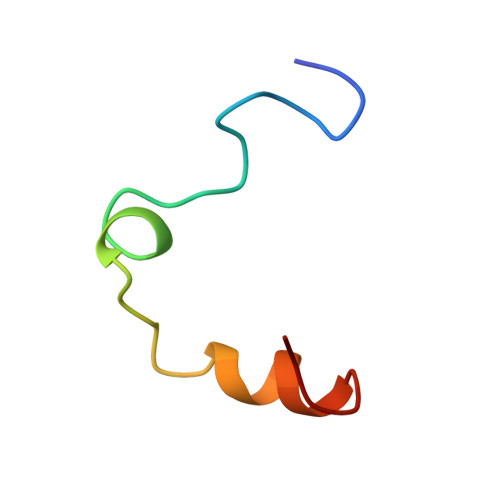
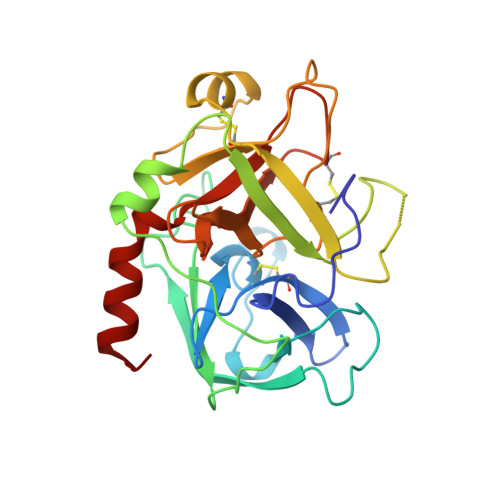
Human alpha-thrombin inhibition by the highly selective compounds N-ethoxycarbonyl-D-Phe-Pro-alpha-azaLys p-nitrophenyl ester and N-carbobenzoxy-Pro-alpha-azaLys p-nitrophenyl ester: a kinetic, thermodynamic and X-ray crystallographic study.
De Simone, G., Balliano, G., Milla, P., Gallina, C., Giordano, C., Tarricone, C., Rizzi, M., Bolognesi, M., Ascenzi, P.(1997) J Mol Biol 269: 558-569
- PubMed: 9217260
- DOI: https://doi.org/10.1006/jmbi.1997.1037
- Primary Citation of Related Structures:
1AE8, 1AFE - PubMed Abstract:
Kinetics, thermodynamics and structural aspects of human alpha-thrombin (thrombin) inhibition by newly synthesized low molecular weight derivatives of alpha-azalysine have been investigated. The thrombin catalyzed hydrolysis of N-ethoxycarbonyl-D-Phe-Pro-alpha-azaLys p-nitrophenyl ester (Eoc-D-Phe-Pro-azaLys-ONp) and N-carbobenzoxy-Pro-alpha-azaLys p-nitrophenyl ester (Cbz-Pro-azaLys-ONp) was investigated at pH 6.2 and 21.0 degrees C, and analyzed in parallel with that of N-alpha-(N,N-dimethylcarbamoyl)-alpha-azalysine p-nitrophenyl ester (Dmc-azaLys-ONp). Decarboxylation following the enzymatic hydrolysis of these p-nitrophenyl esters gave the corresponding 1-peptidyl-2(4-aminobutyl) hydrazines (peptidyl-Abh) showing properties of thrombin competitive inhibitors. Therefore, thermodynamics for the reversible binding of D-Phe-Pro-Abh, Cbz-Pro-Abh and Dmc-Abh to thrombin was examined. These results are consistent with the minimum four-step catalytic mechanism for product inhibition of serine proteinases. Eoc-D-Phe-Pro-azaLys-ONp and Eoc-D-Phe-Pro-Abh display a sub-micromolar affinity for thrombin together with a high selectivity versus homologous plasmatic and pancreatic serine proteinases acting on cationic substrates. The three-dimensional structures of the reversible non-covalent thrombin:Eoc-D-Phe-Pro-Abh and thrombin:Cbz-Pro-Abh complexes have been determined by X-ray crystallography at 2.0 A resolution (R-factor = 0.169 and 0.179, respectively), and analyzed in parallel with that of the thrombin:Dmc-azaLys acyl-enzyme adduct. Both Eoc-D-Phe-Pro-Abh and Cbz-Pro-Abh competitive inhibitors are accommodated in the thrombin active center, spanning the region between the aryl binding site and the S1 primary specificity subsite.
Organizational Affiliation:
Department of Chemistry, University of Napoli, Italy.