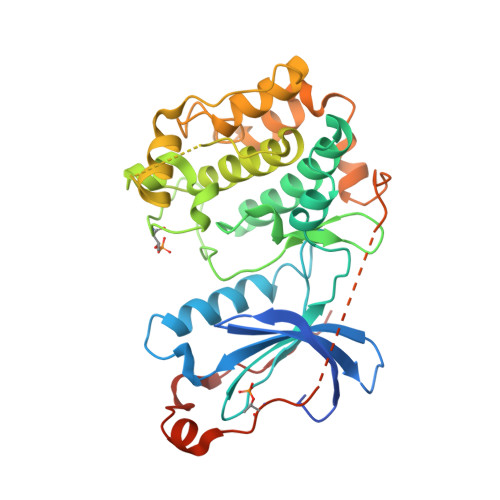
Crystal Structure of the Catalytic Domain of Human Atypical Protein Kinase C-iota Reveals Interaction Mode of Phosphorylation Site in Turn Motif
Messerschmidt, A., Macieira, S., Velarde, M., Baedeker, M., Benda, C., Jestel, A., Brandstetter, H., Neuefeind, T., Blaesse, M.(2005) J Mol Biol 352: 918-931
- PubMed: 16125198
- DOI: https://doi.org/10.1016/j.jmb.2005.07.060
- Primary Citation of Related Structures:
1ZRZ - PubMed Abstract:
Atypical protein kinases C (aPKCs) play critical roles in signaling pathways that control cell growth, differentiation and survival. Therefore, they constitute attractive targets for the development of novel therapeutics against cancer. The crystal structure of the catalytic domain of atypical PKCiota in complex with the bis(indolyl)maleimide inhibitor BIM1 has been determined at 3.0A resolution within the frame of the European Structural Proteomics Project SPINE. The overall structure exhibits the classical bilobal kinase fold and is in its fully activated form. Both phosphorylation sites (Thr403 in the activation loop, and Thr555 in the turn motif) are well defined in the structure and form intramolecular ionic contacts that make an important contribution in stabilizing the active conformation of the catalytic subunit. The phosphorylation site in the hydrophobic motif of atypical PKCs is replaced by the phosphorylation mimic glutamate and this is also clearly seen in the structure of PKCiota (residue 574). This structure determination for the first time provides the architecture of the turn motif phosphorylation site, which is characteristic for PKCs and PKB/AKT, and is completely different from that in PKA. The bound BIM1 inhibitor blocks the ATP-binding site and puts the kinase domain into an intermediate open conformation. The PKCiota-BIM1 complex is the first kinase domain crystal structure of any atypical PKC and constitutes the basis for rational drug design for selective PKCiota inhibitors.
Organizational Affiliation:
Department of Structural Research, Max-Planck-Institute of Biochemistry, Am Klopferspitz 18, 82152 Martinsried, Germany. messersc@biochem.mpg.de